HIV-1 Capsid Rapidly Induces Long-Lived CPSF6 Puncta in Non-Dividing Cells, but Similar Puncta Already Exist in Uninfected T-Cells
- PMID: 38793552
- PMCID: PMC11125723
- DOI: 10.3390/v16050670
HIV-1 Capsid Rapidly Induces Long-Lived CPSF6 Puncta in Non-Dividing Cells, but Similar Puncta Already Exist in Uninfected T-Cells
Abstract
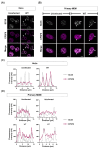
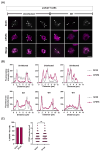
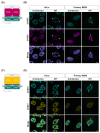
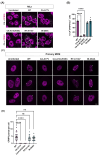
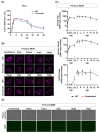
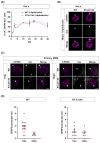
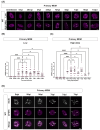
The HIV-1 capsid (CA) protein forms the outer shell of the viral core that is released into the cytoplasm upon infection. CA binds various cellular proteins, including CPSF6, that direct HIV-1 integration into speckle-associated domains in host chromatin. Upon HIV-1 infection, CPSF6 forms puncta in the nucleus. Here, we characterised these CPSF6 puncta further in HeLa cells, T-cells and macrophages and confirmed that integration and reverse transcription are not required for puncta formation. Indeed, we found that puncta formed very rapidly after infection, correlating with the time that CA entered the nucleus. In aphidicolin-treated HeLa cells and macrophages, puncta were detected for the length of the experiment, suggesting that puncta are only lost upon cell division. CA still co-localised with CPSF6 puncta at the latest time points, considerably after the peak of reverse transcription and integration. Intriguingly, the number of puncta induced in macrophages did not correlate with the MOI or the total number of nuclear speckles present in each cell, suggesting that CA/CPSF6 is only directed to a few nuclear speckles. Furthermore, we found that CPSF6 already co-localised with nuclear speckles in uninfected T-cells, suggesting that HIV-1 promotes a natural behaviour of CPSF6.
Keywords: CPSF5; CPSF6; HIV-1; T-cell; capsid; macrophage; nuclear speckle; uncoating.
Conflict of interest statement
The authors declare no conflicts of interest. The sponsors had no role in the design, execution, interpretation, or writing of the study.
Figures
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

