RTCpredictor: identification of read-through chimeric RNAs from RNA sequencing data
- PMID: 38796690
- PMCID: PMC11128028
- DOI: 10.1093/bib/bbae251
RTCpredictor: identification of read-through chimeric RNAs from RNA sequencing data
Abstract
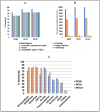
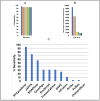
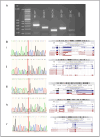
Read-through chimeric RNAs are being recognized as a means to expand the functional transcriptome and contribute to cancer tumorigenesis when mis-regulated. However, current software tools often fail to predict them. We have developed RTCpredictor, utilizing a fast ripgrep tool to search for all possible exon-exon combinations of parental gene pairs. We also added exonic variants allowing searches containing common SNPs. To our knowledge, it is the first read-through chimeric RNA specific prediction method that also provides breakpoint coordinates. Compared with 10 other popular tools, RTCpredictor achieved high sensitivity on a simulated and three real datasets. In addition, RTCpredictor has less memory requirements and faster execution time, making it ideal for applying on large datasets.
Keywords: RNA-Seq; RTCpredictor; chimeric RNA; read-through.
© The Author(s) 2024. Published by Oxford University Press.
Figures


Update of
-
RTCpredictor: Identification of Read-Through Chimeric RNAs from RNA Sequencing Data.bioRxiv [Preprint]. 2023 Feb 3:2023.02.02.526869. doi: 10.1101/2023.02.02.526869. bioRxiv. 2023. Update in: Brief Bioinform. 2024 May 23;25(4):bbae251. doi: 10.1093/bib/bbae251. PMID: 36778443 Free PMC article. Updated. Preprint.

