Deciphering molecular bridges: Unveiling the interplay between metabolic syndrome and Alzheimer's disease through a systems biology approach and drug repurposing
- PMID: 38809924
- PMCID: PMC11135670
- DOI: 10.1371/journal.pone.0304410
Deciphering molecular bridges: Unveiling the interplay between metabolic syndrome and Alzheimer's disease through a systems biology approach and drug repurposing
Abstract
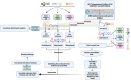
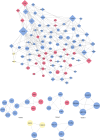
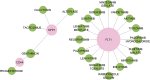
The association between Alzheimer's disease and metabolic disorders as significant risk factors is widely acknowledged. However, the intricate molecular mechanism intertwining these conditions remains elusive. To address this knowledge gap, we conducted a thorough investigation using a bioinformatics method to illuminate the molecular connections and pathways that provide novel perspectives on these disorders' pathological and clinical features. Microarray datasets (GSE5281, GSE122063) from the Gene Expression Omnibus (GEO) database facilitated the way to identify genes with differential expression in Alzheimer's disease (141 genes). Leveraging CoreMine, CTD, and Gene Card databases, we extracted genes associated with metabolic conditions, including hypertension, non-alcoholic fatty liver disease, and diabetes. Subsequent analysis uncovered overlapping genes implicated in metabolic conditions and Alzheimer's disease, revealing shared molecular links. We utilized String and HIPPIE databases to visualize these shared genes' protein-protein interactions (PPI) and constructed a PPI network using Cytoscape and MCODE plugin. SPP1, CD44, IGF1, and FLT1 were identified as crucial molecules in the main cluster of Alzheimer's disease and metabolic syndrome. Enrichment analysis by the DAVID dataset was employed and highlighted the SPP1 as a novel target, with its receptor CD44 playing a significant role in the inflammatory cascade and disruption of insulin signaling, contributing to the neurodegenerative aspects of Alzheimer's disease. ECM-receptor interactions, focal adhesion, and the PI3K/Akt pathways may all mediate these effects. Additionally, we investigated potential medications by repurposing the molecular links using the DGIdb database, revealing Tacrolimus and Calcitonin as promising candidates, particularly since they possess binding sites on the SPP1 molecule. In conclusion, our study unveils crucial molecular bridges between metabolic syndrome and AD, providing insights into their pathophysiology for therapeutic interventions.
Copyright: © 2024 Azizan et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
Identification of crosstalk genes and immune characteristics between Alzheimer's disease and atherosclerosis.Front Immunol. 2024 Aug 12;15:1443464. doi: 10.3389/fimmu.2024.1443464. eCollection 2024. Front Immunol. 2024. PMID: 39188714 Free PMC article.
-
Association between chronic periodontitis and the risk of Alzheimer's disease: combination of text mining and GEO dataset.BMC Oral Health. 2021 Sep 23;21(1):466. doi: 10.1186/s12903-021-01827-2. BMC Oral Health. 2021. PMID: 34556089 Free PMC article.
-
Identification of early Alzheimer's disease subclass and signature genes based on PANoptosis genes.Front Immunol. 2024 Nov 22;15:1462003. doi: 10.3389/fimmu.2024.1462003. eCollection 2024. Front Immunol. 2024. PMID: 39650656 Free PMC article.
-
Common gene signatures and key pathways in hypopharyngeal and esophageal squamous cell carcinoma: Evidence from bioinformatic analysis.Medicine (Baltimore). 2020 Oct 16;99(42):e22434. doi: 10.1097/MD.0000000000022434. Medicine (Baltimore). 2020. PMID: 33080677 Free PMC article.
-
Viral Induced Oxidative and Inflammatory Response in Alzheimer's Disease Pathogenesis with Identification of Potential Drug Candidates: A Systematic Review using Systems Biology Approach.Curr Neuropharmacol. 2019;17(4):352-365. doi: 10.2174/1570159X16666180419124508. Curr Neuropharmacol. 2019. PMID: 29676229 Free PMC article.
Cited by
-
DeepDrug as an expert guided and AI driven drug repurposing methodology for selecting the lead combination of drugs for Alzheimer's disease.Sci Rep. 2025 Jan 15;15(1):2093. doi: 10.1038/s41598-025-85947-7. Sci Rep. 2025. PMID: 39814937 Free PMC article.
-
Osteopontin in Alzheimer's Disease: A Double-Edged Sword in Neurodegeneration and Neuroprotection-A Systematic Review.CNS Neurosci Ther. 2025 Feb;31(2):e70269. doi: 10.1111/cns.70269. CNS Neurosci Ther. 2025. PMID: 39957678 Free PMC article.
-
The Possible Associations between Tauopathies and Atherosclerosis, Diabetes Mellitus, Dyslipidemias, Metabolic Syndrome and Niemann-Pick Disease.Diagnostics (Basel). 2024 Aug 22;14(16):1831. doi: 10.3390/diagnostics14161831. Diagnostics (Basel). 2024. PMID: 39202319 Free PMC article. Review.
-
Linking Metabolic Syndrome to Neurodegeneration Mechanisms and Potential Treatments.Mol Neurobiol. 2025 Apr 24. doi: 10.1007/s12035-025-04947-w. Online ahead of print. Mol Neurobiol. 2025. PMID: 40272771 Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

