A non-canonical role of ELN protects from cellular senescence by limiting iron-dependent regulation of gene expression
- PMID: 38810421
- PMCID: PMC11167390
- DOI: 10.1016/j.redox.2024.103204
A non-canonical role of ELN protects from cellular senescence by limiting iron-dependent regulation of gene expression
Abstract
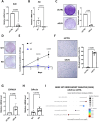
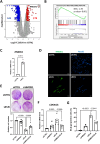
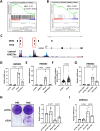
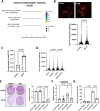
The ELN gene encodes tropoelastin which is used to generate elastic fibers that insure proper tissue elasticity. Decreased amounts of elastic fibers and/or accumulation of bioactive products of their cleavage, named elastokines, are thought to contribute to aging. Cellular senescence, characterized by a stable proliferation arrest and by the senescence-associated secretory phenotype (SASP), increases with aging, fostering the onset and progression of age-related diseases and overall aging, and has so far never been linked with elastin. Here, we identified that decrease in ELN either by siRNA in normal human fibroblasts or by knockout in mouse embryonic fibroblasts results in premature senescence. Surprisingly this effect is independent of elastic fiber degradation or elastokines production, but it relies on the rapid increase in HMOX1 after ELN downregulation. Moreover, the induction of HMOX1 depends on p53 and NRF2 transcription factors, and leads to an increase in iron, further mediating ELN downregulation-induced senescence. Screening of iron-dependent DNA and histones demethylases revealed a role for histone PHF8 demethylase in mediating ELN downregulation-induced senescence. Collectively, these results unveil a role for ELN in protecting cells from cellular senescence through a non-canonical mechanism involving a ROS/HMOX1/iron accumulation/PHF8 histone demethylase pathway reprogramming gene expression towards a senescence program.
Keywords: Cellular senescence; ELN; HMOX1; Iron; PHF8; ROS.
Copyright © 2024 The Authors. Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors have no competing interests to declare.
Figures
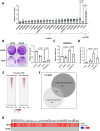
Similar articles
-
ELN regulates cellular senescence: Emerging hypothesis for a non-canonical role.Mech Ageing Dev. 2025 Jun;225:112055. doi: 10.1016/j.mad.2025.112055. Epub 2025 Apr 3. Mech Ageing Dev. 2025. PMID: 40187570 Review.
-
Heme oxygenase-1 affects generation and spontaneous cardiac differentiation of induced pluripotent stem cells.IUBMB Life. 2018 Feb;70(2):129-142. doi: 10.1002/iub.1711. Epub 2018 Jan 9. IUBMB Life. 2018. PMID: 29316264
-
Protective effect of dihydroartemisinin in inhibiting senescence of myeloid-derived suppressor cells from lupus mice via Nrf2/HO-1 pathway.Free Radic Biol Med. 2019 Nov 1;143:260-274. doi: 10.1016/j.freeradbiomed.2019.08.013. Epub 2019 Aug 13. Free Radic Biol Med. 2019. PMID: 31419476
-
Rapamycin inhibits the secretory phenotype of senescent cells by a Nrf2-independent mechanism.Aging Cell. 2017 Jun;16(3):564-574. doi: 10.1111/acel.12587. Epub 2017 Mar 31. Aging Cell. 2017. PMID: 28371119 Free PMC article.
-
p53 Isoforms in Cellular Senescence- and Ageing-Associated Biological and Physiological Functions.Int J Mol Sci. 2019 Nov 29;20(23):6023. doi: 10.3390/ijms20236023. Int J Mol Sci. 2019. PMID: 31795382 Free PMC article. Review.
Cited by
-
PPM1J regulates meat quality feature and glycerophospholipids composition in broiler by modulating protein dephosphorylation.NPJ Sci Food. 2024 Nov 7;8(1):89. doi: 10.1038/s41538-024-00335-1. NPJ Sci Food. 2024. PMID: 39511232 Free PMC article.
-
The Matrix Protein Tropoelastin Prolongs Mesenchymal Stromal Cell Vitality and Delays Senescence During Replicative Aging.Adv Sci (Weinh). 2024 Oct;11(39):e2402168. doi: 10.1002/advs.202402168. Epub 2024 Aug 9. Adv Sci (Weinh). 2024. PMID: 39120048 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

