Escherichia coli self-organizes developmental rosettes
- PMID: 38814871
- PMCID: PMC11161754
- DOI: 10.1073/pnas.2315850121
Escherichia coli self-organizes developmental rosettes
Abstract
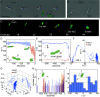
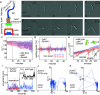
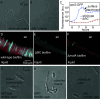
Rosettes are self-organizing, circular multicellular communities that initiate developmental processes, like organogenesis and embryogenesis, in complex organisms. Their formation results from the active repositioning of adhered sister cells and is thought to distinguish multicellular organisms from unicellular ones. Though common in eukaryotes, this multicellular behavior has not been reported in bacteria. In this study, we found that Escherichia coli forms rosettes by active sister-cell repositioning. After division, sister cells "fold" to actively align at the 2- and 4-cell stages of clonal division, thereby producing rosettes with characteristic quatrefoil configuration. Analysis revealed that folding follows an angular random walk, composed of ~1 µm strokes and directional randomization. We further showed that this motion was produced by the flagellum, the extracellular tail whose rotation generates swimming motility. Rosette formation was found to require de novo flagella synthesis suggesting it must balance the opposing forces of Ag43 adhesion and flagellar propulsion. We went on to show that proper rosette formation was required for subsequent morphogenesis of multicellular chains, rpoS gene expression, and formation of hydrostatic clonal-chain biofilms. Moreover, we found self-folding rosette-like communities in the standard motility assay, indicating that this behavior may be a general response to hydrostatic environments in E. coli. These findings establish self-organization of clonal rosettes by a prokaryote and have implications for evolutionary biology, synthetic biology, and medical microbiology.
Keywords: bacteria; development; dynamics; multicellular.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures


References
-
- Knoll A. H., The multiple origins of complex multicellularity. Annu. Rev. Earth Planet. Sci. 39, 217–239 (2011).
-
- Rose C. J., Hammerschmidt K., What do we mean by multicellularity? The evolutionary transitions framework provides answers. Front. Ecol. Evol. 9, 730714 (2021).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

