Comparative chloroplast genomics and phylogenetic analysis of Oreomecon nudicaulis (Papaveraceae)
- PMID: 38816818
- PMCID: PMC11141030
- DOI: 10.1186/s12863-024-01236-8
Comparative chloroplast genomics and phylogenetic analysis of Oreomecon nudicaulis (Papaveraceae)
Abstract
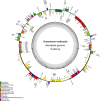
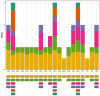
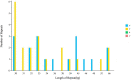
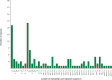
Oreomecon nudicaulis, commonly known as mountain poppy, is a significant perennial herb. In 2022, the species O. nudicaulis, which was previously classified under the genus Papaver, was reclassified within the genus Oreomecon. Nevertheless, the phylogenetic status and chloroplast genome within the genus Oreomecon have not yet been reported. This study elucidates the chloroplast genome sequence and structural features of O. nudicaulis and explores its evolutionary relationships within Papaveraceae. Using Illumina sequencing technology, the chloroplast genome of O. nudicaulis was sequenced, assembled, and annotated. The results indicate that the chloroplast genome of O. nudicaulis exhibits a typical circular quadripartite structure. The chloroplast genome is 153,903 bp in length, with a GC content of 38.87%, containing 84 protein-coding genes, 8 rRNA genes, 38 tRNA genes, and 2 pseudogenes. The genome encodes 25,815 codons, with leucine (Leu) being the most abundant codon, and the most frequently used codon is AUU. Additionally, 129 microsatellite markers were identified, with mononucleotide repeats being the most abundant (53.49%). Our phylogenetic analysis revealed that O. nudicaulis has a relatively close relationship with the genus Meconopsis within the Papaveraceae family. The phylogenetic analysis supported the taxonomic status of O. nudicaulis, as it did not form a clade with other Papaver species, consistent with the revised taxonomy of Papaveraceae. This is the first report of a phylogenomic study of the complete chloroplast genome in the genus Oreomecon, which is a significant genus worldwide. This analysis of the O. nudicaulis chloroplast genome provides a theoretical basis for research on genetic diversity, molecular marker development, and species identification, enriching genetic information and supporting the evolutionary relationships among Papaveraceae.
Keywords: O. nudicaulis; Chloroplast genome; Structural characteristics; Systematic evolution.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
Comparative and Phylogenetic Analysis of the Complete Chloroplast Genomes of Lithocarpus Species (Fagaceae) in South China.Genes (Basel). 2025 May 22;16(6):616. doi: 10.3390/genes16060616. Genes (Basel). 2025. PMID: 40565508 Free PMC article.
-
Comparative analysis of 18 chloroplast genomes reveals genomic diversity and evolutionary dynamics in subtribe Malaxidinae (Orchidaceae).BMC Plant Biol. 2025 Aug 2;25(1):1013. doi: 10.1186/s12870-025-06772-8. BMC Plant Biol. 2025. PMID: 40753185 Free PMC article.
-
Assembly and comparative analysis of the complete mitochondrial genome of Cardiocrinum giganteum: a primitive Liliaceae group with significant scientific research value.BMC Genomics. 2025 Jul 1;26(1):602. doi: 10.1186/s12864-025-11817-1. BMC Genomics. 2025. PMID: 40597596 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of paclitaxel, docetaxel, gemcitabine and vinorelbine in non-small-cell lung cancer.Health Technol Assess. 2001;5(32):1-195. doi: 10.3310/hta5320. Health Technol Assess. 2001. PMID: 12065068
Cited by
-
The Complete Chloroplast Genome of Meconopsis simplicifolia and Its Genetic Comparison to Other Meconopsis Species.Genes (Basel). 2024 Oct 6;15(10):1301. doi: 10.3390/genes15101301. Genes (Basel). 2024. PMID: 39457425 Free PMC article.
References
-
- Editorial Committee of Flora of China . Flora of China. Beijing: Science; 2008. pp. 278–80.
-
- Goldblatt P. Biosystematic studies in Papaver Section Oxytona. Ann Mo Bot Gard. 1974;61:264–96. doi: 10.2307/2395056. - DOI
-
- Christenhusz MJM, Byng JW. The number of known plants species in the World and its annual increase. Phytotaxa. 2016;261(3):201–17. doi: 10.11646/phytotaxa.261.3.1. - DOI
-
- Wei X. The Taxonomic Significance of the Micromorphological and Anatomical Structures of the Taxa in the Family Papaveraceae in Northeastern China [master’s thesis]. Harbin: Harbin Normal University; 2017.
Publication types
MeSH terms
Substances
Grants and funding
- 2022SJYB0090/Philosophy and Social Science Research Project of Jiangsu Province University
- "Public Security Technology" project (2022)/Jiangsu Province "14th Five-Year Plan" Key Construction Discipline
- "Public Security Technology" project (2022)/Jiangsu Province "14th Five-Year Plan" Key Construction Discipline
- "Public Security Technology" project (2022)/Jiangsu Province "14th Five-Year Plan" Key Construction Discipline
- "Public Security Technology" project (2022)/Jiangsu Province "14th Five-Year Plan" Key Construction Discipline
LinkOut - more resources
Full Text Sources
Miscellaneous
