SARS-CoV-2 wastewater variant surveillance: pandemic response leveraging FDA's GenomeTrakr network
- PMID: 38819130
- PMCID: PMC11326115
- DOI: 10.1128/msystems.01415-23
SARS-CoV-2 wastewater variant surveillance: pandemic response leveraging FDA's GenomeTrakr network
Abstract
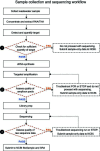
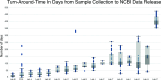
Wastewater surveillance has emerged as a crucial public health tool for population-level pathogen surveillance. Supported by funding from the American Rescue Plan Act of 2021, the FDA's genomic epidemiology program, GenomeTrakr, was leveraged to sequence SARS-CoV-2 from wastewater sites across the United States. This initiative required the evaluation, optimization, development, and publication of new methods and analytical tools spanning sample collection through variant analyses. Version-controlled protocols for each step of the process were developed and published on protocols.io. A custom data analysis tool and a publicly accessible dashboard were built to facilitate real-time visualization of the collected data, focusing on the relative abundance of SARS-CoV-2 variants and sub-lineages across different samples and sites throughout the project. From September 2021 through June 2023, a total of 3,389 wastewater samples were collected, with 2,517 undergoing sequencing and submission to NCBI under the umbrella BioProject, PRJNA757291. Sequence data were released with explicit quality control (QC) tags on all sequence records, communicating our confidence in the quality of data. Variant analysis revealed wide circulation of Delta in the fall of 2021 and captured the sweep of Omicron and subsequent diversification of this lineage through the end of the sampling period. This project successfully achieved two important goals for the FDA's GenomeTrakr program: first, contributing timely genomic data for the SARS-CoV-2 pandemic response, and second, establishing both capacity and best practices for culture-independent, population-level environmental surveillance for other pathogens of interest to the FDA.
Importance: This paper serves two primary objectives. First, it summarizes the genomic and contextual data collected during a Covid-19 pandemic response project, which utilized the FDA's laboratory network, traditionally employed for sequencing foodborne pathogens, for sequencing SARS-CoV-2 from wastewater samples. Second, it outlines best practices for gathering and organizing population-level next generation sequencing (NGS) data collected for culture-free, surveillance of pathogens sourced from environmental samples.
Keywords: FAIR data; GenomeTrakr; SARS-CoV-2; covid-19; data standards; data structures; pathogen genomic surveillance; wastewater based epidemiology; wastewater surveillance.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- World Health Organization . 2021. Weekly epidemiological update - 2 February 2021. Available from: https://www.who.int/publications/m/item/weekly-epidemiological-update---.... Retrieved 24 Nov 2022.
-
- Stevens EL, Carleton HA, Beal J, Tillman GE, Lindsey RL, Lauer AC, Pightling A, Jarvis KG, Ottesen A, Ramachandran P, et al. 2022. The use of whole-genome sequencing by the Federal interagency collaboration for genomics for food and feed safety in the United States. J Food Prot 85:755–772. doi: 10.4315/JFP-21-437 - DOI - PubMed
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

