Development of a genome-scale metabolic model for the lager hybrid yeast S. pastorianus to understand the evolution of metabolic pathways in industrial settings
- PMID: 38819150
- PMCID: PMC11237392
- DOI: 10.1128/msystems.00429-24
Development of a genome-scale metabolic model for the lager hybrid yeast S. pastorianus to understand the evolution of metabolic pathways in industrial settings
Abstract
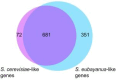
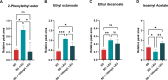
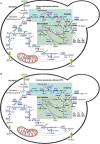
In silico tools such as genome-scale metabolic models have shown to be powerful for metabolic engineering of microorganisms. Saccharomyces pastorianus is a complex aneuploid hybrid between the mesophilic Saccharomyces cerevisiae and the cold-tolerant Saccharomyces eubayanus. This species is of biotechnological importance because it is the primary yeast used in lager beer fermentation and is also a key model for studying the evolution of hybrid genomes, including expression pattern of ortholog genes, composition of protein complexes, and phenotypic plasticity. Here, we created the iSP_1513 GSMM for S. pastorianus CBS1513 to allow top-down computational approaches to predict the evolution of metabolic pathways and to aid strain optimization in production processes. The iSP_1513 comprises 4,062 reactions, 1,808 alleles, and 2,747 metabolites, and takes into account the functional redundancy in the gene-protein-reaction rule caused by the presence of orthologous genes. Moreover, a universal algorithm to constrain GSMM reactions using transcriptome data was developed as a python library and enabled the integration of temperature as parameter. Essentiality data sets, growth data on various carbohydrates and volatile metabolites secretion were used to validate the model and showed the potential of media engineering to improve specific flavor compounds. The iSP_1513 also highlighted the different contributions of the parental sub-genomes to the oxidative and non-oxidative parts of the pentose phosphate pathway. Overall, the iSP_1513 GSMM represent an important step toward understanding the metabolic capabilities, evolutionary trajectories, and adaptation potential of S. pastorianus in different industrial settings.
Importance: Genome-scale metabolic models (GSMM) have been successfully applied to predict cellular behavior and design cell factories in several model organisms, but no models to date are currently available for hybrid species due to their more complex genetics and general lack of molecular data. In this study, we generated a bespoke GSMM, iSP_1513, for this industrial aneuploid hybrid Saccharomyces pastorianus, which takes into account the aneuploidy and functional redundancy from orthologous parental alleles. This model will (i) help understand the metabolic capabilities and adaptive potential of S. pastorianus (domestication processes), (ii) aid top-down predictions for strain development (industrial biotechnology), and (iii) allow predictions of evolutionary trajectories of metabolic pathways in aneuploid hybrids (evolutionary genetics).
Keywords: genome-scale metabolic model; yeast hybrid.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Himalayan Saccharomyces eubayanus Genome Sequences Reveal Genetic Markers Explaining Heterotic Maltotriose Consumption by Saccharomyces pastorianus Hybrids.Appl Environ Microbiol. 2019 Oct 30;85(22):e01516-19. doi: 10.1128/AEM.01516-19. Print 2019 Nov 15. Appl Environ Microbiol. 2019. PMID: 31519660 Free PMC article.
-
Industrially Applicable De Novo Lager Yeast Hybrids with a Unique Genomic Architecture: Creation and Characterization.Appl Environ Microbiol. 2021 Jan 15;87(3):e02434-20. doi: 10.1128/AEM.02434-20. Print 2021 Jan 15. Appl Environ Microbiol. 2021. PMID: 33188002 Free PMC article.
-
Lager-brewing yeasts in the era of modern genetics.FEMS Yeast Res. 2019 Nov 1;19(7):foz063. doi: 10.1093/femsyr/foz063. FEMS Yeast Res. 2019. PMID: 31553794 Free PMC article. Review.
-
Saccharomyces pastorianus: genomic insights inspiring innovation for industry.Yeast. 2015 Jan;32(1):17-27. doi: 10.1002/yea.3033. Epub 2014 Sep 23. Yeast. 2015. PMID: 25088523 Review.
-
Transcriptional Profile of the Industrial Hybrid Saccharomyces pastorianus Reveals Temperature-Dependent Allele Expression Bias and Preferential Orthologous Protein Assemblies.Mol Biol Evol. 2021 Dec 9;38(12):5437-5452. doi: 10.1093/molbev/msab282. Mol Biol Evol. 2021. PMID: 34550394 Free PMC article.
Cited by
-
Impact of inter-species hybridisation on antifungal drug response in the Saccharomyces genus.BMC Genomics. 2024 Dec 2;25(1):1165. doi: 10.1186/s12864-024-11009-3. BMC Genomics. 2024. PMID: 39623390 Free PMC article.
-
Flux Sampling Suggests Metabolic Signatures of High Antibody-Producing CHO Cells.Biotechnol Bioeng. 2025 Jul;122(7):1898-1913. doi: 10.1002/bit.28982. Epub 2025 Apr 11. Biotechnol Bioeng. 2025. PMID: 40219633 Free PMC article.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

