Structural conversion of the spidroin C-terminal domain during assembly of spider silk fibers
- PMID: 38821983
- PMCID: PMC11143275
- DOI: 10.1038/s41467-024-49111-5
Structural conversion of the spidroin C-terminal domain during assembly of spider silk fibers
Abstract
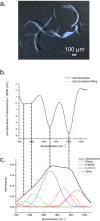
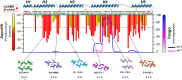
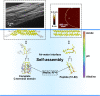
The major ampullate Spidroin 1 (MaSp1) is the main protein of the dragline spider silk. The C-terminal (CT) domain of MaSp1 is crucial for the self-assembly into fibers but the details of how it contributes to the fiber formation remain unsolved. Here we exploit the fact that the CT domain can form silk-like fibers by itself to gain knowledge about this transition. Structural investigations of fibers from recombinantly produced CT domain from E. australis MaSp1 reveal an α-helix to β-sheet transition upon fiber formation and highlight the helix No4 segment as most likely to initiate the structural conversion. This prediction is corroborated by the finding that a peptide corresponding to helix No4 has the ability of pH-induced conversion into β-sheets and self-assembly into nanofibrils. Our results provide structural information about the CT domain in fiber form and clues about its role in triggering the structural conversion of spidroins during fiber assembly.
© 2024. The Author(s).
Conflict of interest statement
M.H. has shares in the research company Spiber Technologies AB. L.G. is employed by the research company Spiber Technologies AB. The remaining authors declare no competing interests.
Figures




Similar articles
-
Recombinant Production, Characterization, and Fiber Spinning of an Engineered Short Major Ampullate Spidroin (MaSp1s).Biomacromolecules. 2017 Apr 10;18(4):1365-1372. doi: 10.1021/acs.biomac.7b00090. Epub 2017 Mar 6. Biomacromolecules. 2017. PMID: 28233980
-
Nearly complete 1H, 13C and 15N chemical shift assignment of monomeric form of N-terminal domain of Nephila clavipes major ampullate spidroin 2.Biomol NMR Assign. 2020 Oct;14(2):335-338. doi: 10.1007/s12104-020-09972-5. Epub 2020 Aug 6. Biomol NMR Assign. 2020. PMID: 32767002
-
NMR assignment and dynamics of the dimeric form of soluble C-terminal domain major ampullate spidroin 2 from Latrodectus hesperus.Biomol NMR Assign. 2023 Dec;17(2):249-255. doi: 10.1007/s12104-023-10150-6. Epub 2023 Sep 5. Biomol NMR Assign. 2023. PMID: 37668860
-
Complexity of Spider Dragline Silk.Biomacromolecules. 2022 May 9;23(5):1827-1840. doi: 10.1021/acs.biomac.1c01682. Epub 2022 Apr 4. Biomacromolecules. 2022. PMID: 35378031 Free PMC article. Review.
-
Silk Spinning in Silkworms and Spiders.Int J Mol Sci. 2016 Aug 9;17(8):1290. doi: 10.3390/ijms17081290. Int J Mol Sci. 2016. PMID: 27517908 Free PMC article. Review.
Cited by
-
Silk Protein Gene Engineering and Its Applications: Recent Advances in Biomedicine Driven by Molecular Biotechnology.Drug Des Devel Ther. 2025 Jan 25;19:599-626. doi: 10.2147/DDDT.S504783. eCollection 2025. Drug Des Devel Ther. 2025. PMID: 39881670 Free PMC article. Review.
-
Recombinant production of spider silk protein in Physcomitrella photobioreactors.Plant Cell Rep. 2025 Apr 26;44(5):103. doi: 10.1007/s00299-025-03485-y. Plant Cell Rep. 2025. PMID: 40287554 Free PMC article.
-
Progress in Multiscale Modeling of Silk Materials.Biomacromolecules. 2024 Nov 11;25(11):6987-7014. doi: 10.1021/acs.biomac.4c01122. Epub 2024 Oct 22. Biomacromolecules. 2024. PMID: 39438248 Free PMC article. Review.
References
-
- Vollrath F, Barth P, Basedow A, Engström W, List H. Local tolerance to spider silks and protein polymers in vivo. Vivo. 2002;16:229–234. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

