Development of an miRFP680-Based Fluorescent Calcium Ion Biosensor Using End-Optimized Transposons
- PMID: 38822813
- PMCID: PMC11218748
- DOI: 10.1021/acssensors.4c00727
Development of an miRFP680-Based Fluorescent Calcium Ion Biosensor Using End-Optimized Transposons
Abstract
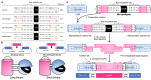
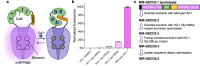
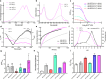
The development of new or improved single fluorescent protein (FP)-based biosensors (SFPBs), particularly those with excitation and emission at near-infrared wavelengths, is important for the continued advancement of biological imaging applications. In an effort to accelerate the development of new SFPBs, we report modified transposons for the transposase-based creation of libraries of FPs randomly inserted into analyte binding domains, or vice versa. These modified transposons feature ends that are optimized to minimize the length of the linkers that connect the FP to the analyte binding domain. We rationalized that shorter linkers between the domains should result in more effective allosteric coupling between the analyte binding-dependent conformational change in the binding domain and the fluorescence modulation of the chromophore of the FP domain. As a proof of concept, we employed end-modified Mu transposons for the discovery of SFPB prototypes based on the insertion of two circularly permuted red FPs (mApple and FusionRed) into binding proteins for l-lactate and spermidine. Using an analogous approach, we discovered calcium ion (Ca2+)-specific SFPBs by random insertion of calmodulin (CaM)-RS20 into miRFP680, a particularly bright near-infrared (NIR) FP based on a biliverdin (BV)-binding fluorescent protein. Starting from an miRFP680-based Ca2+ biosensor prototype, we performed extensive directed evolution, including under BV-deficient conditions, to create highly optimized biosensors designated the NIR-GECO3 series. We have extensively characterized the NIR-GECO3 series and explored their utility for biological Ca2+ imaging. The methods described in this work will serve to accelerate SFPB development and open avenues for further exploration and optimization of SFPBs across a spectrum of biological applications.
Keywords: biliverdin-binding fluorescent protein; cell signaling; directed evolution; fluorescence microscopy; heme oxygenase; protein engineering; transposons.
Conflict of interest statement
The authors declare no competing financial interest.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

