A functional variant rs10409772 in FUT6 promoter regulates colorectal cancer progression through PKA/CREB signaling
- PMID: 38823257
- PMCID: PMC11176829
- DOI: 10.1016/j.tranon.2024.102011
A functional variant rs10409772 in FUT6 promoter regulates colorectal cancer progression through PKA/CREB signaling
Abstract
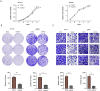
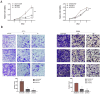
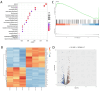
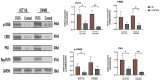
Fucosyltransferase 6 (FUT6) is overexpressed in colorectal cancer tissue according to TCGA samples and immunohistochemistry results of a tissue microarray. FUT6 effects cell migration, tumor formation and proliferation of colorectal cancer cells in different essays. FUT6 promotes cancer cell proliferation in vitro and colorectal tumorigenesis in vivo by upregulating PKA/CREB pathway activation. Moreover, FUT6 expression is regulated by rs10409772 shown in the luciferase essays, a single nucleotide polymorphism in the promoter of FUT6. Our study suggests that elevated expression of FUT6 promotes PKA/CREB signaling, which in turn augments colorectal carcinogenesis, indicating a potential therapeutic target for colorectal cancer patients with increased FUT6 expression.
Keywords: Colorectal cancer; FUT6; PKA/CREB signaling; Single nucleotide polymorphism.
Copyright © 2024. Published by Elsevier Inc.
Conflict of interest statement
Declaration of competing interest There is no competing interest among authors.
Figures




Similar articles
-
miR-125a-3p/FUT5-FUT6 axis mediates colorectal cancer cell proliferation, migration, invasion and pathological angiogenesis via PI3K-Akt pathway.Cell Death Dis. 2017 Aug 3;8(8):e2968. doi: 10.1038/cddis.2017.352. Cell Death Dis. 2017. PMID: 28771224 Free PMC article.
-
FUT6 inhibits the proliferation, migration, invasion, and EGF-induced EMT of head and neck squamous cell carcinoma (HNSCC) by regulating EGFR/ERK/STAT signaling pathway.Cancer Gene Ther. 2023 Jan;30(1):182-191. doi: 10.1038/s41417-022-00530-w. Epub 2022 Sep 23. Cancer Gene Ther. 2023. PMID: 36151332
-
MicroRNA-106b targets FUT6 to promote cell migration, invasion, and proliferation in human breast cancer.IUBMB Life. 2016 Sep;68(9):764-75. doi: 10.1002/iub.1541. Epub 2016 Aug 13. IUBMB Life. 2016. PMID: 27519168
-
Molecular genetics of alpha-L-fucosyltransferase genes (H, Se, Le, FUT4, FUT5 and FUT6).Transfus Clin Biol. 1994;1(2):91-7. doi: 10.1016/s1246-7820(94)80002-2. Transfus Clin Biol. 1994. PMID: 8019653 Review.
-
Complex roles of cAMP-PKA-CREB signaling in cancer.Exp Hematol Oncol. 2020 Nov 24;9(1):32. doi: 10.1186/s40164-020-00191-1. Exp Hematol Oncol. 2020. PMID: 33292604 Free PMC article. Review.
References
-
- Wong C.C., Yu J. Gut microbiota in colorectal cancer development and therapy. Nat. Rev. Clin. Oncol. 2023;20(7):429–452. [J] - PubMed
-
- Sharma B., Angurana S., Bhat A., et al. Genetic analysis of colorectal carcinoma using high throughput single nucleotide polymorphism genotyping technique within the population of Jammu and Kashmir. Mol. Biol. Rep. 2021;48(8):5889–5895. [J] - PubMed
LinkOut - more resources
Full Text Sources

