Diverse genotypes of norovirus genogroup I and II contamination in environmental water in Thailand during the COVID-19 outbreak from 2020 to 2022
- PMID: 38823781
- PMCID: PMC11401460
- DOI: 10.1016/j.virs.2024.05.010
Diverse genotypes of norovirus genogroup I and II contamination in environmental water in Thailand during the COVID-19 outbreak from 2020 to 2022
Abstract
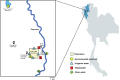
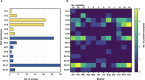
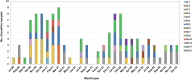
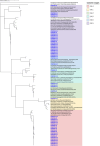
Noroviruses (NoVs) are the most significant viral pathogens associated with waterborne and foodborne outbreaks of nonbacterial acute gastroenteritis in humans worldwide. This study aimed to investigate the prevalence and diversity of NoVs contaminated in the environmental water in Chiang Mai, Thailand. A total of 600 environmental water samples were collected from ten sampling sites in Chiang Mai from July 2020 to December 2022. The presence of NoV genogroups I (GI), GII, and GIV were examined using real-time RT-PCR assay. The genotype of the virus was determined by nucleotide sequencing and phylogenetic analysis. The results showed that NoV GI and GII were detected at 8.5% (51/600) and 11.7% (70/600) of the samples tested, respectively. However, NoV GIV was not detected in this study. NoV circulated throughout the year, with a higher detection rate during the winter season. Six NoV GI genotypes (GI.1-GI.6) and eight NoV GII genotypes (GII.2, GII.3, GII.7, GII.8, GII.10, GII.13, GII.17, and GII.21) were identified. Among 121 NoV strains detected, GII.17 was the most predominant genotype (24.8%, 30 strains), followed by GII.2 (21.5%, 26 strains), GI.3 (17.4%, 21 strains), and GI.4 (16.5%, 20 strains). Notably, NoV GII.3, GII.7, GII.8, and GII.10 were detected for the first time in water samples in this area. This study provides insight into the occurrence and seasonal pattern of NoV along with novel findings of NoV strains in environmental water in Thailand during the COVID-19 outbreak. Our findings emphasize the importance of further surveillance studies to monitor viral contamination in environmental water.
Keywords: Detection; Environmental water; Norovirus (NoVs); Thailand; Wastewater.
Copyright © 2024 The Authors. Publishing services by Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors declare no conflict of interest relevant to this article.
Figures
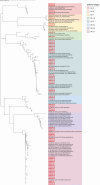
Similar articles
-
Genetic diversity of norovirus genogroup I, II, IV and sapovirus in environmental water in Thailand.J Infect Public Health. 2020 Oct;13(10):1481-1489. doi: 10.1016/j.jiph.2020.05.004. Epub 2020 May 31. J Infect Public Health. 2020. PMID: 32493670
-
Emergence of new norovirus variants and genetic heterogeneity of noroviruses and sapoviruses in children admitted to hospital with diarrhea in Thailand.J Med Virol. 2010 Feb;82(2):289-96. doi: 10.1002/jmv.21640. J Med Virol. 2010. PMID: 20029815
-
Novel norovirus recombinants and of GII.4 sub-lineages associated with outbreaks between 2006 and 2010 in Belgium.Virol J. 2011 Jun 18;8:310. doi: 10.1186/1743-422X-8-310. Virol J. 2011. PMID: 21682917 Free PMC article.
-
Molecular epidemiology and genotype distributions of noroviruses and sapoviruses in Thailand 2000-2016: A review.J Med Virol. 2018 Apr;90(4):617-624. doi: 10.1002/jmv.25019. Epub 2018 Jan 23. J Med Virol. 2018. PMID: 29315631 Review.
-
The epidemiology of published norovirus outbreaks: a review of risk factors associated with attack rate and genogroup.Epidemiol Infect. 2012 Jul;140(7):1161-72. doi: 10.1017/S0950268812000234. Epub 2012 Mar 26. Epidemiol Infect. 2012. PMID: 22444943 Free PMC article.
Cited by
-
The genetic diversity of genogroup I noroviruses causing acute gastroenteritis outbreaks in Beijing between 2014 and 2023.Heliyon. 2024 Oct 10;10(20):e39202. doi: 10.1016/j.heliyon.2024.e39202. eCollection 2024 Oct 30. Heliyon. 2024. PMID: 39640656 Free PMC article.
-
Quantification and Genotyping of Norovirus in Aerosols from Wastewater Treatment Plants in Thailand.Food Environ Virol. 2025 May 23;17(2):31. doi: 10.1007/s12560-025-09647-1. Food Environ Virol. 2025. PMID: 40407958 Free PMC article.
References
-
- Boonchan M., Motomura K., Inoue K., Ode H., Chu P.Y., Lin M., Iwatani Y., Ruchusatsawat K., Guntapong R., Tacharoenmuang R., Chantaroj S., Tatsumi M., Takeda N., Sangkitporn S. Distribution of norovirus genotypes and subtypes in river water by ultra-deep sequencing-based analysis. Lett. Appl. Microbiol. 2017;65:98–104. - PubMed
-
- Cannon J.L., Barclay L., Collins N.R., Wikswo M.E., Castro C.J., Magana L.C., Gregoricus N., Marine R.L., Chhabra P., Vinjé J. Genetic and epidemiologic trends of norovirus outbreaks in the United States from 2013 to 2016 demonstrated emergence of novel GII.4 recombinant viruses. J. Clin. Microbiol. 2017;55:2208–2221. - PMC - PubMed
-
- Douglas A., Sandmann F.G., Allen D.J., Celma C.C., Beard S., Larkin L. Impact of COVID-19 on national surveillance of norovirus in England and potential risk of increased disease activity in 2021. J. Hosp. Infect. 2021;112:124–126. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

