This is a preprint.
The impact of chronic pain on brain gene expression
- PMID: 38826319
- PMCID: PMC11142271
- DOI: 10.1101/2024.05.20.24307630
The impact of chronic pain on brain gene expression
Update in
-
The impact of chronic pain on brain gene expression.Pain. 2025 Dec 1;166(12):e689-e702. doi: 10.1097/j.pain.0000000000003707. Epub 2025 Jul 7. Pain. 2025. PMID: 40623285 Free PMC article.
Abstract
Background: Chronic pain affects one fifth of American adults, contributing significant public health burden. Chronic pain mechanisms can be further understood through investigating brain gene expression.
Methods: We tested differentially expressed genes (DEGs) in chronic pain, migraine, lifetime fentanyl and oxymorphone use, and with chronic pain genetic risk in four brain regions (dACC, DLPFC, MeA, BLA) and imputed cell type expression data from 304 postmortem donors. We compared findings across traits and with independent transcriptomics resources, and performed gene-set enrichment.
Results: We identified two chronic pain DEGs: B4GALT and VEGFB in bulk dACC. We found over 2000 (primarily BLA microglia) chronic pain cell type DEGs. Findings were enriched for mouse microglia pain genes, and for hypoxia and immune response. Cross-trait DEG overlap was minimal.
Conclusions: Chronic pain-associated gene expression is heterogeneous across cell type, largely distinct from that in pain-related traits, and shows BLA microglia are a key cell type.
Figures

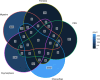
References
-
- Yong R. J., Mullins P. M. & Bhattacharyya N. Prevalence of chronic pain among adults in the United States. PAIN 163, e328 (2022). - PubMed
-
- Terminology | International Association for the Study of Pain. International Association for the Study of Pain (IASP) https://www.iasp-pain.org/resources/terminology/.
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
