This is a preprint.
Genome-wide nucleosome and transcription factor responses to genetic perturbations reveal chromatin-mediated mechanisms of transcriptional regulation
- PMID: 38826400
- PMCID: PMC11142231
- DOI: 10.1101/2024.05.24.595391
Genome-wide nucleosome and transcription factor responses to genetic perturbations reveal chromatin-mediated mechanisms of transcriptional regulation
Abstract
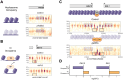
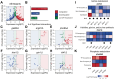
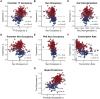
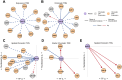
Epigenetic mechanisms contribute to gene regulation by altering chromatin accessibility through changes in transcription factor (TF) and nucleosome occupancy throughout the genome. Despite numerous studies focusing on changes in gene expression, the intricate chromatin-mediated regulatory code remains largely unexplored on a comprehensive scale. We address this by employing a factor-agnostic, reverse-genetics approach that uses MNase-seq to capture genome-wide TF and nucleosome occupancies in response to the individual deletion of 201 transcriptional regulators in Saccharomyces cerevisiae, thereby assaying nearly one million mutant-gene interactions. We develop a principled approach to identify and quantify chromatin changes genome-wide, observing differences in TF and nucleosome occupancy that recapitulate well-established pathways identified by gene expression data. We also discover distinct chromatin signatures associated with the up- and downregulation of genes, and use these signatures to reveal regulatory mechanisms previously unexplored in expression-based studies. Finally, we demonstrate that chromatin features are predictive of transcriptional activity and leverage these features to reconstruct chromatin-based transcriptional regulatory networks. Overall, these results illustrate the power of an approach combining genetic perturbation with high-resolution epigenomic profiling; the latter enables a close examination of the interplay between TFs and nucleosomes genome-wide, providing a deeper, more mechanistic understanding of the complex relationship between chromatin organization and transcription.
Figures



Similar articles
-
Differential effects of chromatin regulators and transcription factors on gene regulation: a nucleosomal perspective.Bioinformatics. 2011 Jan 15;27(2):147-52. doi: 10.1093/bioinformatics/btq637. Epub 2010 Nov 11. Bioinformatics. 2011. PMID: 21075748
-
In vivo effects of histone H3 depletion on nucleosome occupancy and position in Saccharomyces cerevisiae.PLoS Genet. 2012;8(6):e1002771. doi: 10.1371/journal.pgen.1002771. Epub 2012 Jun 21. PLoS Genet. 2012. PMID: 22737086 Free PMC article.
-
RoboCOP: jointly computing chromatin occupancy profiles for numerous factors from chromatin accessibility data.Nucleic Acids Res. 2021 Aug 20;49(14):7925-7938. doi: 10.1093/nar/gkab553. Nucleic Acids Res. 2021. PMID: 34255854 Free PMC article.
-
Changing the DNA landscape: putting a SPN on chromatin.Curr Top Microbiol Immunol. 2003;274:171-201. doi: 10.1007/978-3-642-55747-7_7. Curr Top Microbiol Immunol. 2003. PMID: 12596908 Review.
-
Nucleosome-Omics: A Perspective on the Epigenetic Code and 3D Genome Landscape.Genes (Basel). 2022 Jun 22;13(7):1114. doi: 10.3390/genes13071114. Genes (Basel). 2022. PMID: 35885897 Free PMC article. Review.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
