Chromosome-level genome assemblies of 2 hemichordates provide new insights into deuterostome origin and chromosome evolution
- PMID: 38829909
- PMCID: PMC11175523
- DOI: 10.1371/journal.pbio.3002661
Chromosome-level genome assemblies of 2 hemichordates provide new insights into deuterostome origin and chromosome evolution
Abstract
Deuterostomes are a monophyletic group of animals that includes Hemichordata, Echinodermata (together called Ambulacraria), and Chordata. The diversity of deuterostome body plans has made it challenging to reconstruct their ancestral condition and to decipher the genetic changes that drove the diversification of deuterostome lineages. Here, we generate chromosome-level genome assemblies of 2 hemichordate species, Ptychodera flava and Schizocardium californicum, and use comparative genomic approaches to infer the chromosomal architecture of the deuterostome common ancestor and delineate lineage-specific chromosomal modifications. We show that hemichordate chromosomes (1N = 23) exhibit remarkable chromosome-scale macrosynteny when compared to other deuterostomes and can be derived from 24 deuterostome ancestral linkage groups (ALGs). These deuterostome ALGs in turn match previously inferred bilaterian ALGs, consistent with a relatively short transition from the last common bilaterian ancestor to the origin of deuterostomes. Based on this deuterostome ALG complement, we deduced chromosomal rearrangement events that occurred in different lineages. For example, a fusion-with-mixing event produced an Ambulacraria-specific ALG that subsequently split into 2 chromosomes in extant hemichordates, while this homologous ALG further fused with another chromosome in sea urchins. Orthologous genes distributed in these rearranged chromosomes are enriched for functions in various developmental processes. We found that the deeply conserved Hox clusters are located in highly rearranged chromosomes and that maintenance of the clusters are likely due to lower densities of transposable elements within the clusters. We also provide evidence that the deuterostome-specific pharyngeal gene cluster was established via the combination of 3 pre-assembled microsyntenic blocks. We suggest that since chromosomal rearrangement events and formation of new gene clusters may change the regulatory controls of developmental genes, these events may have contributed to the evolution of diverse body plans among deuterostomes.
Copyright: © 2024 Lin et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
D.S.R. is the paid consultant and shareholder of Dovetail Genomics. The other authors have declared that no competing interests exist.
Figures





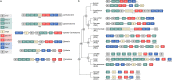
References
-
- Satoh N. Chordate Origins and Evolution: The Molecular Evolutionary Road to Vertebrates. Chordate Origins and Evolution: The Molecular Evolutionary Road to Vertebrates. 2016:1–206. WOS:000404599900015.
MeSH terms
LinkOut - more resources
Full Text Sources

