Reduced protein-coding transcript diversity in severe dengue emphasises the role of alternative splicing
- PMID: 38830771
- PMCID: PMC11147948
- DOI: 10.26508/lsa.202402683
Reduced protein-coding transcript diversity in severe dengue emphasises the role of alternative splicing
Erratum in
-
Correction: Reduced protein-coding transcript diversity in severe dengue emphasises the role of alternative splicing.Life Sci Alliance. 2024 Jun 14;7(9):e202402882. doi: 10.26508/lsa.202402882. Print 2024 Sep. Life Sci Alliance. 2024. PMID: 38876798 Free PMC article.
Abstract
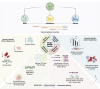
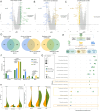
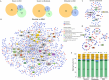
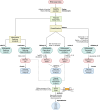
Dengue fever, a neglected tropical arboviral disease, has emerged as a global health concern in the past decade. Necessitating a nuanced comprehension of the intricate dynamics of host-virus interactions influencing disease severity, we analysed transcriptomic patterns using bulk RNA-seq from 112 age- and gender-matched NS1 antigen-confirmed hospital-admitted dengue patients with varying severity. Severe cases exhibited reduced platelet count, increased lymphocytosis, and neutropenia, indicating a dysregulated immune response. Using bulk RNA-seq, our analysis revealed a minimal overlap between the differentially expressed gene and transcript isoform, with a distinct expression pattern across the disease severity. Severe patients showed enrichment in retained intron and nonsense-mediated decay transcript biotypes, suggesting altered splicing efficiency. Furthermore, an up-regulated programmed cell death, a haemolytic response, and an impaired interferon and antiviral response at the transcript level were observed. We also identified the potential involvement of the RBM39 gene among others in the innate immune response during dengue viral pathogenesis, warranting further investigation. These findings provide valuable insights into potential therapeutic targets, underscoring the importance of exploring transcriptomic landscapes between different disease sub-phenotypes in infectious diseases.
© 2024 Mehta et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures






Similar articles
-
Dengue virus targets RBM10 deregulating host cell splicing and innate immune response.Nucleic Acids Res. 2020 Jul 9;48(12):6824-6838. doi: 10.1093/nar/gkaa340. Nucleic Acids Res. 2020. PMID: 32432721 Free PMC article.
-
Immune and vascular modulation by HERVs: the role of CXCR1 and IL18RAP in dengue severity progression.Front Immunol. 2025 Mar 7;16:1557588. doi: 10.3389/fimmu.2025.1557588. eCollection 2025. Front Immunol. 2025. PMID: 40124360 Free PMC article.
-
Unraveling potential gene biomarkers for dengue infection through RNA sequencing.Virus Genes. 2025 Feb;61(1):26-37. doi: 10.1007/s11262-024-02114-2. Epub 2024 Oct 14. Virus Genes. 2025. PMID: 39397194 Free PMC article.
-
Host Genetic Polymorphisms Influencing Susceptibility to Dengue.DNA Cell Biol. 2018 Oct;37(10):805-807. doi: 10.1089/dna.2018.4372. Epub 2018 Aug 16. DNA Cell Biol. 2018. PMID: 30113225 Review.
-
DENGUE VIRUS VIRULENCE AND DISEASES SEVERITY.Southeast Asian J Trop Med Public Health. 2015;46 Suppl 1:35-42. Southeast Asian J Trop Med Public Health. 2015. PMID: 26506730 Review.
Cited by
-
Exploiting host kinases to combat dengue virus infection and disease.Antiviral Res. 2025 Sep;241:106172. doi: 10.1016/j.antiviral.2025.106172. Epub 2025 May 8. Antiviral Res. 2025. PMID: 40348023 Review.
-
Suboptimal dengue genome leverages non-canonical translation mechanisms.iScience. 2025 Apr 15;28(5):112428. doi: 10.1016/j.isci.2025.112428. eCollection 2025 May 16. iScience. 2025. PMID: 40641555 Free PMC article.
-
Genomic hotspots in the DENV-2 serotype (E, NS4B, and NS5 genes) are associated with dengue disease severity in the endemic region of India.PLoS Negl Trop Dis. 2025 Apr 29;19(4):e0013034. doi: 10.1371/journal.pntd.0013034. eCollection 2025 Apr. PLoS Negl Trop Dis. 2025. PMID: 40299925 Free PMC article.
References
-
- Dengue and Severe Dengue (2024). Available at: https://www.who.int/news-room/fact-sheets/detail/dengue-and-severe-dengue.
-
- DENGUE SITUATION IN ::INDIA National Center for Vector Borne Diseases Control (NCVBDC) (2024). Available at: https://ncvbdc.mohfw.gov.in/index4.php?lang=1&level=0&linkid=431&lid=3715.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
