Screening and functional verification of drought resistance-related genes in castor bean seeds
- PMID: 38831288
- PMCID: PMC11145773
- DOI: 10.1186/s12870-024-04997-7
Screening and functional verification of drought resistance-related genes in castor bean seeds
Abstract
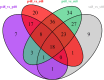
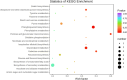
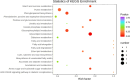
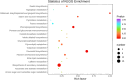
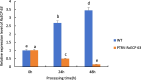
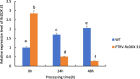
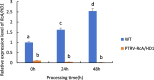
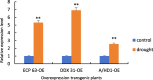
Drought is one of the natural stresses that greatly impact plants. Castor bean (Ricinus communis L.) is an oil crop with high economic value. Drought is one of the factors limiting castor bean growth. The drought resistance mechanisms of castor bean have become a research focus. In this study, we used castor germinating embryos as experimental materials, and screened genes related to drought resistance through physiological measurements, proteomics and metabolomics joint analysis; castor drought-related genes were subjected to transient silencing expression analysis in castor leaves to validate their drought-resistant functions, and heterologous overexpression and backward complementary expression in Arabidopsis thaliana, and analysed the mechanism of the genes' response to the participation of Arabidopsis thaliana in drought-resistance.Three drought tolerance-related genes, RcECP 63, RcDDX 31 and RcA/HD1, were obtained by screening and analysis, and transient silencing of expression in castor leaves further verified that these three genes corresponded to drought stress, and heterologous overexpression and back-complementary expression of the three genes in Arabidopsis thaliana revealed that the function of these three genes in drought stress response.In this study, three drought tolerance related genes, RcECP 63, RcDDX 31 and RcA/HD1, were screened and analysed for gene function, which were found to be responsive to drought stress and to function in drought stress, laying the foundation for the study of drought tolerance mechanism in castor bean.
Keywords: RcA/HD1; RcDDX31; RcECP63; Castor bean; Drought stress.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
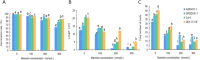
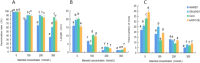
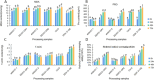
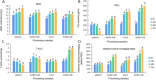
Figures









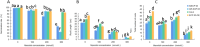


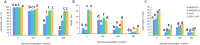


Similar articles
-
Genome-wide identification of castor bean class III peroxidase genes and analysis of expression patterns under abiotic stresses.BMC Plant Biol. 2025 Jul 11;25(1):900. doi: 10.1186/s12870-025-06945-5. BMC Plant Biol. 2025. PMID: 40646456 Free PMC article.
-
Genome-wide identification of core components of ABA signaling and transcriptome analysis reveals gene circuits involved in castor bean (Ricinus communis L.) response to drought.Gene. 2023 Oct 20;883:147668. doi: 10.1016/j.gene.2023.147668. Epub 2023 Jul 25. Gene. 2023. PMID: 37500024
-
Genome-wide analysis of LOG family genes in castor and RcLOG5 enhances drought, salt, and cold stress tolerance in Arabidopsis thaliana.Gene. 2024 Jun 30;913:148398. doi: 10.1016/j.gene.2024.148398. Epub 2024 Mar 20. Gene. 2024. PMID: 38518901
-
Identification and characterization of the RcTCP gene family and its expression in response to abiotic stresses in castor bean.BMC Genomics. 2024 Jul 4;25(1):670. doi: 10.1186/s12864-024-10347-6. BMC Genomics. 2024. PMID: 38965476 Free PMC article.
-
Identification of reference genes for quantitative RT-PCR analysis of microRNAs and mRNAs in castor bean (Ricinus communis L.) under drought stress.Plant Physiol Biochem. 2016 Sep;106:101-7. doi: 10.1016/j.plaphy.2016.02.024. Epub 2016 Mar 19. Plant Physiol Biochem. 2016. PMID: 27156134
Cited by
-
Study on the physiological mechanism and molecular regulatory network of Blumea balsamifera in response to drought stress.BMC Plant Biol. 2025 Jul 2;25(1):803. doi: 10.1186/s12870-025-06916-w. BMC Plant Biol. 2025. PMID: 40604416 Free PMC article.
References
-
- Li Z, Li HH, Yu SK. Sl WDR204 gene positively regulates drought stress in tomato plants. Anhui Agric Sci. 2019;47(1):96–98.
-
- Toker C, Canci H, Yildirim T. Evaluation of perennial wild Cicer species for drought resistance. Genet Resour Crop Evol. 2007;54(8):1781–1786. doi: 10.1007/s10722-006-9197-y. - DOI
-
- Yao Q. Study on the mechanism of drought stress signalling response of Arabidopsis AtRACK1 protein. Yangzhou University. 2007.
-
- Zhao JD. Effective ways to improve drought resistance of plants. Livestock Feed Sci. 2009; 30(2):50-51+117.
MeSH terms
Substances
Grants and funding
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 2021BS03036/Analysis of the R2R3-MYB transcription factor gene family in castor bean and functional resolution of RcMYB61 in response to drought stress
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 31860071/National Natural Science Foundation of China
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2020114/Ministry of Education New Agricultural Research and Reform Practice Project
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2021MS03008/Upper Level Project of Inner Mongolia Natural Science Foundation
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 2022/Inner Mongolia Autonomous Region Grassland Talent Innovation Team - Rolling Support Programme for Castor Molecular Breeding Research Innovation Talent Team
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 21082/2021 Research Project on Higher Education Teaching Reform of the National People's Committee
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- 237/Inner Mongolia University for Nationalities 2022 Basic Research Operating Expenses of Colleges and Universities directly under the Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
- MDK2021011, MDK2022014/Project of the Open Fund of the Collaborative Innovation Centre for Castor Industry in Inner Mongolia Autonomous Region
LinkOut - more resources
Full Text Sources

