Development of a rapamycin-inducible protein-knockdown system in the unicellular red alga Cyanidioschyzon merolae
- PMID: 38833589
- PMCID: PMC11376382
- DOI: 10.1093/plphys/kiae316
Development of a rapamycin-inducible protein-knockdown system in the unicellular red alga Cyanidioschyzon merolae
Abstract
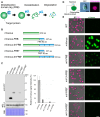
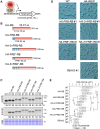
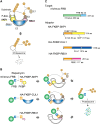
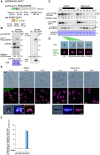
An inducible protein-knockdown system is highly effective for investigating the functions of proteins and mechanisms essential for the survival and growth of organisms. However, this technique is not available in photosynthetic eukaryotes. The unicellular red alga Cyanidioschyzon merolae possesses a very simple cellular and genomic architecture and is genetically tractable but lacks RNA interference machinery. In this study, we developed a protein-knockdown system in this alga. The constitutive system utilizes the destabilizing activity of the FK506-binding protein 12 (FKBP12)-rapamycin-binding (FRB) domain of human target of rapamycin kinase or its derivatives to knock down target proteins. In the inducible system, rapamycin treatment induces the heterodimerization of the human FRB domain fused to the target proteins with the human FKBP fused to S-phase kinase-associated protein 1 or Cullin 1, subunits of the SCF E3 ubiquitin ligase. This results in the rapid degradation of the target proteins through the ubiquitin-proteasome pathway. With this system, we successfully degraded endogenous essential proteins such as the chloroplast division protein dynamin-related protein 5B and E2 transcription factor, a regulator of the G1/S transition, within 2 to 3 h after rapamycin administration, enabling the assessment of resulting phenotypes. This rapamycin-inducible protein-knockdown system contributes to the functional analysis of genes whose disruption leads to lethality.
© The Author(s) 2024. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Conflict of interest statement
Conflict of interest statement. None declared.
Figures

Similar articles
-
Expression of budding yeast FKBP12 confers rapamycin susceptibility to the unicellular red alga Cyanidioschyzon merolae.Biochem Biophys Res Commun. 2013 Sep 20;439(2):264-9. doi: 10.1016/j.bbrc.2013.08.045. Epub 2013 Aug 22. Biochem Biophys Res Commun. 2013. PMID: 23973485
-
Construction of a rapamycin-susceptible strain of the unicellular red alga Cyanidioschyzon merolae for analysis of the target of rapamycin (TOR) function.J Gen Appl Microbiol. 2017 Nov 17;63(5):305-309. doi: 10.2323/jgam.2017.02.002. Epub 2017 Sep 26. J Gen Appl Microbiol. 2017. PMID: 28954966 No abstract available.
-
Target of rapamycin (TOR) plays a critical role in triacylglycerol accumulation in microalgae.Plant Mol Biol. 2015 Oct;89(3):309-18. doi: 10.1007/s11103-015-0370-6. Epub 2015 Sep 8. Plant Mol Biol. 2015. PMID: 26350402
-
The Unicellular Red Alga Cyanidioschyzon merolae-The Simplest Model of a Photosynthetic Eukaryote.Plant Cell Physiol. 2021 Sep 30;62(6):926-941. doi: 10.1093/pcp/pcab052. Plant Cell Physiol. 2021. PMID: 33836072 Free PMC article. Review.
-
Halcyon days of TOR: Reflections on the multiple independent discovery of the yeast and mammalian TOR proteins.Gene. 2019 Apr 15;692:145-155. doi: 10.1016/j.gene.2018.12.046. Epub 2019 Jan 9. Gene. 2019. PMID: 30639424 Review.
Cited by
-
Auxin promotes chloroplast division by increasing the expression of chloroplast division genes.Plant Cell Rep. 2024 Dec 31;44(1):20. doi: 10.1007/s00299-024-03415-4. Plant Cell Rep. 2024. PMID: 39741196
-
Manipulation of targeted protein degradation in plant biology.Biochem Soc Trans. 2025 Apr 9;53(2):409-18. doi: 10.1042/BST20230939. Biochem Soc Trans. 2025. PMID: 40209052 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

