The NRC0 gene cluster of sensor and helper NLR immune receptors is functionally conserved across asterid plants
- PMID: 38833594
- PMCID: PMC11371149
- DOI: 10.1093/plcell/koae154
The NRC0 gene cluster of sensor and helper NLR immune receptors is functionally conserved across asterid plants
Abstract
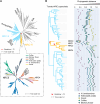
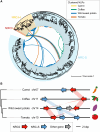
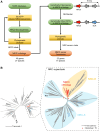
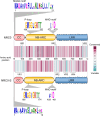
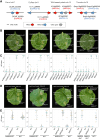
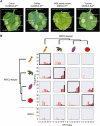
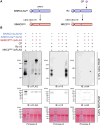
Nucleotide-binding domain and leucine-rich repeat-containing receptor (NLR) proteins can form complex receptor networks to confer innate immunity. An NLR-REQUIRED FOR CELL DEATH (NRC) is a phylogenetically related node that functions downstream of a massively expanded network of disease resistance proteins that protect against multiple plant pathogens. In this study, we used phylogenomic methods to reconstruct the macroevolution of the NRC family. One of the NRCs, termed NRC0, is the only family member shared across asterid plants, leading us to investigate its evolutionary history and genetic organization. In several asterid species, NRC0 is genetically clustered with other NLRs that are phylogenetically related to NRC-dependent disease resistance genes. This prompted us to hypothesize that the ancestral state of the NRC network is an NLR helper-sensor gene cluster that was present early during asterid evolution. We provide support for this hypothesis by demonstrating that NRC0 is essential for the hypersensitive cell death that is induced by its genetically linked sensor NLR partners in 4 divergent asterid species: tomato (Solanum lycopersicum), wild sweet potato (Ipomoea trifida), coffee (Coffea canephora), and carrot (Daucus carota). In addition, activation of a sensor NLR leads to higher-order complex formation of its genetically linked NRC0, similar to other NRCs. Our findings map out contrasting evolutionary dynamics in the macroevolution of the NRC network over the last 125 million years, from a functionally conserved NLR gene cluster to a massive genetically dispersed network.
© The Author(s) 2024. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Conflict of interest statement
Conflict of interest statement. S.K. receives funding from industry on NLR biology and is a cofounder of start-up companies that focus on plant disease resistance. M.P.C. and S.K. have filed patents on NLR biology. M.P.C. has received fees from Resurrect Bio Ltd, a start-up company related to NLR biology.
Figures


Similar articles
-
NRC Immune receptor networks show diversified hierarchical genetic architecture across plant lineages.Plant Cell. 2024 Sep 3;36(9):3399-3418. doi: 10.1093/plcell/koae179. Plant Cell. 2024. PMID: 38922300 Free PMC article.
-
A root-specific NLR network mediates immune signaling of resistance genes against plant parasitic nematodes.Plant Cell. 2025 Jul 1;37(7):koaf145. doi: 10.1093/plcell/koaf145. Plant Cell. 2025. PMID: 40581623 Free PMC article.
-
An atypical NLR protein modulates the NRC immune receptor network in Nicotiana benthamiana.PLoS Genet. 2023 Jan 19;19(1):e1010500. doi: 10.1371/journal.pgen.1010500. eCollection 2023 Jan. PLoS Genet. 2023. PMID: 36656829 Free PMC article.
-
NLR receptors in plant immunity: making sense of the alphabet soup.EMBO Rep. 2023 Oct 9;24(10):e57495. doi: 10.15252/embr.202357495. Epub 2023 Aug 21. EMBO Rep. 2023. PMID: 37602936 Free PMC article. Review.
-
Activation and Regulation of NLR Immune Receptor Networks.Plant Cell Physiol. 2022 Oct 31;63(10):1366-1377. doi: 10.1093/pcp/pcac116. Plant Cell Physiol. 2022. PMID: 35941738 Review.
Cited by
-
Birth and expansion of NRC immune receptors across the largest group of flowering plants.Plant Cell. 2024 Sep 3;36(9):3320-3321. doi: 10.1093/plcell/koae185. Plant Cell. 2024. PMID: 38917235 Free PMC article. No abstract available.
-
NRC Immune receptor networks show diversified hierarchical genetic architecture across plant lineages.Plant Cell. 2024 Sep 3;36(9):3399-3418. doi: 10.1093/plcell/koae179. Plant Cell. 2024. PMID: 38922300 Free PMC article.
-
Subfunctionalization of NRC3 altered the genetic structure of the Nicotiana NRC network.PLoS Genet. 2024 Sep 12;20(9):e1011402. doi: 10.1371/journal.pgen.1011402. eCollection 2024 Sep. PLoS Genet. 2024. PMID: 39264953 Free PMC article.
-
A root-specific NLR network mediates immune signaling of resistance genes against plant parasitic nematodes.Plant Cell. 2025 Jul 1;37(7):koaf145. doi: 10.1093/plcell/koaf145. Plant Cell. 2025. PMID: 40581623 Free PMC article.
-
Helper NLR immune protein NRC3 evolved to evade inhibition by a cyst nematode virulence effector.PLoS Genet. 2025 Apr 9;21(4):e1011653. doi: 10.1371/journal.pgen.1011653. eCollection 2025 Apr. PLoS Genet. 2025. PMID: 40202957 Free PMC article.
References
-
- Ahn HK, Lin X, Olave-Achury AC, Derevnina L, Contreras MP, Kourelis J, Wu CH, Kamoun S, Jones JDG. Effector-dependent activation and oligomerization of plant NRC class helper NLRs by sensor NLR immune receptors Rpi-amr3 and Rpi-amr1. EMBO J. 2023:42(5):e111484. 10.15252/embj.2022111484 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

