TRIM71 mutations cause a neurodevelopmental syndrome featuring ventriculomegaly and hydrocephalus
- PMID: 38833623
- PMCID: PMC11629693
- DOI: 10.1093/brain/awae175
TRIM71 mutations cause a neurodevelopmental syndrome featuring ventriculomegaly and hydrocephalus
Abstract
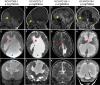
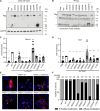
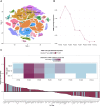
Congenital hydrocephalus, characterized by cerebral ventriculomegaly, is one of the most common reasons for paediatric brain surgery. Recent studies have implicated lin-41 (lineage variant 41)/TRIM71 (tripartite motif 71) as a candidate congenital hydrocephalus risk gene; however, TRIM71 variants have not been systematically examined in a large patient cohort or conclusively linked with an OMIM syndrome. Through cross-sectional analysis of the largest assembled cohort of patients with cerebral ventriculomegaly, including neurosurgically-treated congenital hydrocephalus (totalling 2697 parent-proband trios and 8091 total exomes), we identified 13 protein-altering de novo variants (DNVs) in TRIM71 in unrelated children exhibiting variable ventriculomegaly, congenital hydrocephalus, developmental delay, dysmorphic features and other structural brain defects, including corpus callosum dysgenesis and white matter hypoplasia. Eight unrelated patients were found to harbour arginine variants, including two recurrent missense DNVs, at homologous positions in RPXGV motifs of different NHL domains. Seven patients with rare, damaging, unphased or transmitted variants of uncertain significance were also identified. NHL-domain variants of TRIM71 exhibited impaired binding to the canonical TRIM71 target CDKN1A; other variants failed to direct the subcellular localization of TRIM71 to processing bodies. Single-cell transcriptomic analysis of human embryos revealed expression of TRIM71 in early first-trimester neural stem cells of the brain. These data show TRIM71 is essential for human brain morphogenesis and that TRIM71 mutations cause a novel neurodevelopmental syndrome that we term 'TRIM71-associated developmental disorders (TADD)', featuring variable ventriculomegaly, congenital hydrocephalus and other structural brain defects.
Keywords: de novo variants; TRIM71; brain development; hydrocephalus; neural stem cells; structural brain disorders.
© The Author(s) 2024. Published by Oxford University Press on behalf of the Guarantors of Brain. All rights reserved. For commercial re-use, please contact reprints@oup.com for reprints and translation rights for reprints. All other permissions can be obtained through our RightsLink service via the Permissions link on the article page on our site—for further information please contact journals.permissions@oup.com.
Conflict of interest statement
The authors report no competing interests.
Figures

References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

