Isocucurbitacin B inhibits glioma growth through PI3K/AKT pathways and increases glioma sensitivity to TMZ by inhibiting hsa-mir-1286a
- PMID: 38835342
- PMCID: PMC11149100
- DOI: 10.20517/cdr.2024.01
Isocucurbitacin B inhibits glioma growth through PI3K/AKT pathways and increases glioma sensitivity to TMZ by inhibiting hsa-mir-1286a
Abstract
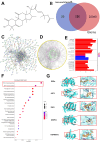
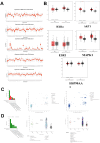
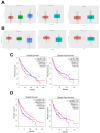
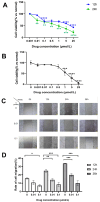
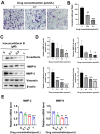
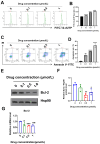
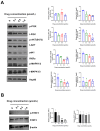
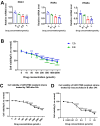
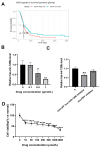
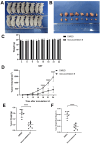
Aim: Glioma accounts for 81% of all cancers of the nervous system cancers and presents one of the most drug-resistant malignancies, resulting in a relatively high mortality rate. Despite extensive efforts, the complete treatment options for glioma remain elusive. The effect of isocucurbitacin B (isocuB), a natural compound extracted from melon pedicels, on glioma has not been investigated. This study aims to investigate the inhibitory effect of isocuB on glioma and elucidate its underlying mechanisms, with the objective of developing it as a potential therapeutic agent for glioma. Methods: We used network pharmacology and bioinformatics analysis to predict potential targets and associated pathways of isocuB in glioma. Subsequently, the inhibitory effect of isocuB on glioma and its related mechanisms were assessed through Counting Kit-8 (CCK-8), wound healing, transwell, Western blot (WB), reverse transcription-quantitative polymerase chain reaction (RT-qPCR), and other in vitro experiments, alongside tumor formation experiments in nude mice. Results: Based on this investigation, it suggested that isocuB might inhibit the growth of gliomas through the PI3K-AKT and MAPK pathways. Additionally, we proposed that isocuB may enhance glioma drug sensitivity to temozolomide (TMZ) via modulation of hsa-mir-1286a. The CCK-8 assay revealed that isocuB exhibited inhibitory effects on U251 and U87 proliferation and outperformed TMZ. Wound healing and transwell experiments showed that isocuB inhibited the invasion and migration of U251 cells by suppressing the activity of MMP-2/9, N-cadherin, and Vimentin. The TdT-mediated dUTP-biotin nick end labeling (TUNEL) and flow cytometry (FCM) assays revealed that isocuB induced cell apoptosis through inhibition of BCL-2. Subsequently, we conducted RT-qPCR and WB experiments, which revealed that PI3K/AKT and MAPK pathways might be involved in the mechanism of the inhibition isocuB on glioma. Additionally, isocuB promoted the sensitivity of glioma U251 to TMZ by inhibiting hsa-mir-1286a. Furthermore, we constructed TMZ-resistant U251 strains and demonstrated effective inhibition by isocuB against these resistant strains. Finally, we confirmed that isocuB can inhibit tumor growth in vivo through experiments on tumors in nude mice. Conclusion: IsocuB may protect against glioma by acting on the PI3K/AKT and MAPK pathways and promote the sensitivity of glioma U251 to TMZ by inhibiting hsa-mir-1286a.
Keywords: Isocucurbitacin B; drug sensitivity; glioma; hsa-mir-1286a; network pharmacology.
© The Author(s) 2024.
Conflict of interest statement
All authors declared that there are no conflicts of interest.
Figures
References
LinkOut - more resources
Full Text Sources
Research Materials
