A novel mutation in the OTOF gene in a Chinese family with auditory neuropathy
- PMID: 38836175
- PMCID: PMC11145404
- DOI: 10.5582/irdr.2024.01004
A novel mutation in the OTOF gene in a Chinese family with auditory neuropathy
Abstract
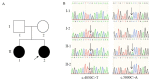
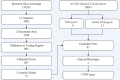
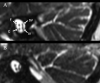
Gene therapy for monogenic auditory neuropathy (AN) has successfully improved hearing function in target gene-deficient mice. Accurate genetic diagnosis can not only clarify the etiology but also accurately locate the lesion site, providing a basis for gene therapy and guiding patient intervention and management strategies. In this study, we collected data from a family with a pair of sisters with prelingual deafness. According to their auditory tests, subject Ⅱ-1 was diagnosed with profound sensorineural hearing loss (SNHL), Ⅱ-2 was diagnosed with AN, Ⅰ-1 was diagnosed with high-frequency SNHL, and Ⅰ-2 had normal hearing. Using whole-exome sequencing (WES), one nonsense mutation, c.4030C>T (p.R1344X), and one missense mutation, c.5000C>A (p.A1667D), in the OTOF (NM_001287489.1) gene were identified in the two siblings. Their parents were heterozygous carriers of c.5000C>A (father) and c.4030C>T (mother). We hypothesized that c.5000C>A is a novel pathogenic mutation. Thus, subject Ⅱ-1 should also be diagnosed with AN caused by OTOF mutations. These findings not only expand the OTOF gene mutation spectrum for AN but also indicate that WES is an effective approach for accurately diagnosing AN.
Keywords: OTOF gene; auditory neuropathy; whole-exome sequencing.
2024, International Research and Cooperation Association for Bio & Socio - Sciences Advancement.
Conflict of interest statement
The authors have no conflicts of interest to disclose.
Figures


Similar articles
-
Novel compound heterozygous mutations in the OTOF Gene identified by whole-exome sequencing in auditory neuropathy spectrum disorder.BMC Med Genet. 2017 Mar 23;18(1):35. doi: 10.1186/s12881-017-0400-0. BMC Med Genet. 2017. PMID: 28335750 Free PMC article.
-
A multicenter study on the prevalence and spectrum of mutations in the otoferlin gene (OTOF) in subjects with nonsyndromic hearing impairment and auditory neuropathy.Hum Mutat. 2008 Jun;29(6):823-31. doi: 10.1002/humu.20708. Hum Mutat. 2008. PMID: 18381613
-
Novel OTOF gene mutations identified using a massively parallel DNA sequencing technique in DFNB9 deafness.Acta Otolaryngol. 2018 Oct;138(10):865-870. doi: 10.1080/00016489.2018.1476777. Epub 2018 Aug 3. Acta Otolaryngol. 2018. PMID: 30073893
-
Targeted next generation sequencing reveals OTOF mutations in auditory neuropathy spectrum disorder.Int J Pediatr Otorhinolaryngol. 2018 Dec;115:19-23. doi: 10.1016/j.ijporl.2018.09.008. Epub 2018 Sep 14. Int J Pediatr Otorhinolaryngol. 2018. PMID: 30368385 Review.
-
Cochlear Implantation Outcomes in Patients With OTOF Mutations.Front Neurosci. 2020 May 21;14:447. doi: 10.3389/fnins.2020.00447. eCollection 2020. Front Neurosci. 2020. PMID: 32508568 Free PMC article. Review.
Cited by
-
Pathogenesis and research progress of OTOF gene related auditory neuropathy: a retrospective review.Am J Transl Res. 2025 Mar 15;17(3):1643-1650. doi: 10.62347/JDLC8070. eCollection 2025. Am J Transl Res. 2025. PMID: 40226018 Free PMC article. Review.
References
-
- Chinese Multi-center Research Collaborative Group on Clinical Diagnosis and Intervention of Auditory Neuropathy; Editorial Board of Chinese Journal of Otorhinolaryngology Head and Neck Surgery; Society of Otorhinolaryngology Head and Neck Surgery, Chinese Medical Association, China Division; International Association of Physicians in Audiology, Society of Audiology and Vestibular Medicine; China International Exchange and Promotive Association for Medical and Health Care. Chinese clinical practice guideline of auditory neuropathy (version 2022). Zhonghua Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 2022; 57:241-262. (in Chinese) - PubMed
-
- Manchaiah VK, Zhao F, Danesh AA, Duprey R. The genetic basis of auditory neuropathy spectrum disorder (ANSD). Int J Pediatr Otorhinolaryngol. 2011; 75:151-158. - PubMed
-
- Schoen CJ, Emery SB, Thorne MC, Ammana HR, Sliwerska E, Arnett J, Hortsch M, Hannan F, Burmeister M, Lesperance MM. Increased activity of Diaphanous homolog 3 (DIAPH3)/diaphanous causes hearing defects in humans with auditory neuropathy and in Drosophila. Proc Natl Acad Sci USA. 2010; 107:13396-13401. - PMC - PubMed
-
- Moser T, Starr A. Auditory neuropathy - neural and synaptic mechanisms. Nat Rev Neurol. 2016; 12:135-149. - PubMed

