Image-based discrimination of the early stages of mesenchymal stem cell differentiation
- PMID: 38837346
- PMCID: PMC11321037
- DOI: 10.1091/mbc.E24-02-0095
Image-based discrimination of the early stages of mesenchymal stem cell differentiation
Abstract
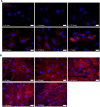
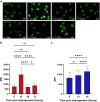
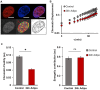
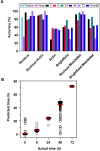
Mesenchymal stem cells (MSCs) are self-renewing, multipotent cells, which can be used in cellular and tissue therapeutics. MSCs cell number can be expanded in vitro, but premature differentiation results in reduced cell number and compromised therapeutic efficacies. Current techniques fail to discriminate the "stem-like" population from early stages (12 h) of differentiated MSC population. Here, we imaged nuclear structure and actin architecture using immunofluorescence and used deep learning-based computer vision technology to discriminate the early stages (6-12 h) of MSC differentiation. Convolutional neural network models trained by nucleus and actin images have high accuracy in reporting MSC differentiation; nuclear images alone can identify early stages of differentiation. Concurrently, we show that chromatin fluidity and heterochromatin levels or localization change during early MSC differentiation. This study quantifies changes in cell architecture during early MSC differentiation and describes a novel image-based diagnostic tool that could be widely used in MSC culture, expansion and utilization.
Conflict of interest statement
Conflicts of interests: The authors declare no financial conflict of interest.
Figures

References
-
- Anghileri E, Marconi S, Pignatelli A, Cifelli P, Galié M, Sbarbati A, Krampera M, Belluzzi O, Bonetti B (2008). Neuronal differentiation potential of human adipose-derived mesenchymal stem cells. Stem Cells Dev 17, 909–916. - PubMed
-
- Boland MV, Markey MK, Murphy RF (1998). Automated recognition of patterns characteristic of subcellular structures in fluorescence microscopy images. Cytometry 33, 366–375 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

