AlphaKnot 2.0: a web server for the visualization of proteins' knotting and a database of knotted AlphaFold-predicted models
- PMID: 38842945
- PMCID: PMC11223836
- DOI: 10.1093/nar/gkae443
AlphaKnot 2.0: a web server for the visualization of proteins' knotting and a database of knotted AlphaFold-predicted models
Abstract
The availability of 3D protein models is rapidly increasing with the development of structure prediction algorithms. With the expanding availability of data, new ways of analysis, especially topological analysis, of those predictions are becoming necessary. Here, we present the updated version of the AlphaKnot service that provides a straightforward way of analyzing structure topology. It was designed specifically to determine knot types of the predicted structure models, however, it can be used for all structures, including the ones solved experimentally. AlphaKnot 2.0 provides the user's ability to obtain the knowledge necessary to assess the topological correctness of the model. Both probabilistic and deterministic knot detection methods are available, together with various visualizations (including a trajectory of simplification steps to highlight the topological complexities). Moreover, the web server provides a list of proteins similar to the queried model within AlphaKnot's database and returns their knot types for direct comparison. We pre-calculated the topology of high-quality models from the AlphaFold Database (4th version) and there are now more than 680.000 knotted models available in the AlphaKnot database. AlphaKnot 2.0 is available at https://alphaknot.cent.uw.edu.pl/.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


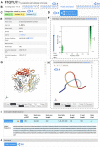
References
-
- Lin Z., Akin H., Rao R., Hie B., Zhu Z., Lu W., Smetanin N., Verkuil R., Kabeli O., Shmueli Y. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science. 2023; 379:1123–1130. - PubMed
-
- Varadi M., Anyango S., Deshpande M., Nair S., Natassia C., Yordanova G., Yuan D., Stroe O., Wood G., Laydon A. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 2022; 50:D439–D444. - PMC - PubMed
-
- Sulkowska J.I. On folding of entangled proteins: knots, lassos, links and -curves. Curr. Opin. Struc. Biol. 2020; 60:131–141. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

