High-throughput screening unveils nitazoxanide as a potent PRRSV inhibitor by targeting NMRAL1
- PMID: 38844461
- PMCID: PMC11156899
- DOI: 10.1038/s41467-024-48807-y
High-throughput screening unveils nitazoxanide as a potent PRRSV inhibitor by targeting NMRAL1
Abstract
Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) poses a major threat to the global swine industry, yet effective prevention and control measures remain elusive. This study unveils Nitazoxanide (NTZ) as a potent inhibitor of PRRSV both in vitro and in vivo. Through High-Throughput Screening techniques, 16 potential anti-PRRSV compounds are identified from a library comprising FDA-approved and pharmacopeial drugs. We show that NTZ displays strong efficacy in reducing PRRSV proliferation and transmission in a swine model, alleviating viremia and lung damage. Additionally, Tizoxanide (TIZ), the primary metabolite of NTZ, has been identified as a facilitator of NMRAL1 dimerization. This finding potentially sheds light on the underlying mechanism contributing to TIZ's role in augmenting the sensitivity of the IFN-β pathway. These results indicate the promising potential of NTZ as a repurposed therapeutic agent for Porcine Reproductive and Respiratory Syndrome (PRRS). Additionally, they provide valuable insights into the antiviral mechanisms underlying NTZ's effectiveness.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



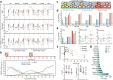


References
-
- Keffaber, K. K. Reproductive failure of unknown etiology. Am. Assoc. Swine Pract. Newsl.1, 1–9 (1989).
-
- Baron T, et al. Report on the first outbreaks of the porcine reproductive and respiratory syndrome (PRRS) in France. Diagnosis and viral isolation. Ann. Rech. Vet. 1992;23:161–166. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

