Single-cell transcriptomic atlas reveals increased regeneration in diseased human inner ear balance organs
- PMID: 38844821
- PMCID: PMC11156867
- DOI: 10.1038/s41467-024-48491-y
Single-cell transcriptomic atlas reveals increased regeneration in diseased human inner ear balance organs
Abstract
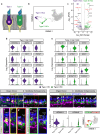
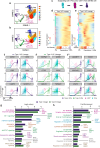
Mammalian inner ear hair cell loss leads to permanent hearing and balance dysfunction. In contrast to the cochlea, vestibular hair cells of the murine utricle have some regenerative capacity. Whether human utricular hair cells regenerate in vivo remains unknown. Here we procured live, mature utricles from organ donors and vestibular schwannoma patients, and present a validated single-cell transcriptomic atlas at unprecedented resolution. We describe markers of 13 sensory and non-sensory cell types, with partial overlap and correlation between transcriptomes of human and mouse hair cells and supporting cells. We further uncover transcriptomes unique to hair cell precursors, which are unexpectedly 14-fold more abundant in vestibular schwannoma utricles, demonstrating the existence of ongoing regeneration in humans. Lastly, supporting cell-to-hair cell trajectory analysis revealed 5 distinct patterns of dynamic gene expression and associated pathways, including Wnt and IGF-1 signaling. Our dataset constitutes a foundational resource, accessible via a web-based interface, serving to advance knowledge of the normal and diseased human inner ear.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





References
MeSH terms
Substances
Grants and funding
- R01 AG081608/AG/NIA NIH HHS/United States
- R01 DC016318/DC/NIDCD NIH HHS/United States
- K08 DC019683/DC/NIDCD NIH HHS/United States
- T32 DC015209/DC/NIDCD NIH HHS/United States
- U24 DC015910/DC/NIDCD NIH HHS/United States
- UL1 TR001085/TR/NCATS NIH HHS/United States
- R01 DC021110/DC/NIDCD NIH HHS/United States
- S10 OD018220/OD/NIH HHS/United States
- R01 DC020879/DC/NIDCD NIH HHS/United States
- UL1 TR003142/TR/NCATS NIH HHS/United States
- R01 DC016409/DC/NIDCD NIH HHS/United States
- F30 DC015698/DC/NIDCD NIH HHS/United States
- U24 DC020857/DC/NIDCD NIH HHS/United States
- R21 DC015879/DC/NIDCD NIH HHS/United States
- R21 DC022058/DC/NIDCD NIH HHS/United States
- R21 DC019457/DC/NIDCD NIH HHS/United States
- S10 OD021763/OD/NIH HHS/United States
- R01 DC016919/DC/NIDCD NIH HHS/United States
- R01 DC013910/DC/NIDCD NIH HHS/United States
- K24 DC020986/DC/NIDCD NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

