Subtractive proteomics-based vaccine targets annotation and reverse vaccinology approaches to identify multiepitope vaccine against Plesiomonas shigelloides
- PMID: 38845922
- PMCID: PMC11153098
- DOI: 10.1016/j.heliyon.2024.e31304
Subtractive proteomics-based vaccine targets annotation and reverse vaccinology approaches to identify multiepitope vaccine against Plesiomonas shigelloides
Abstract
Plesiomonas shigelloides, an aquatic bacterium belonging to the Enterobacteriaceae family, is a frequent cause of gastroenteritis with diarrhea and gastrointestinal severe disease. Despite decades of research, discovering a licensed and globally accessible vaccine is still years away. Developing a putative vaccine that can combat the Plesiomonas shigelloides infection by boosting population immunity against P. shigelloides is direly needed. In the framework of the current study, the entire proteome of P. shigelloides was explored using subtractive genomics integrated with the immunoinformatics approach for designing an effective vaccine construct against P. shigelloides. The overall stability of the vaccine construct was evaluated using molecular docking, which demonstrated that MEV showed higher binding affinities with toll-like receptors (TLR4: 51.5 ± 10.3, TLR2: 60.5 ± 9.2) and MHC receptors(MHCI: 79.7 ± 11.2 kcal/mol, MHCII: 70.4 ± 23.7). Further, the therapeutic efficacy of the vaccine construct for generating an efficient immune response was evaluated by computational immunological simulation. Finally, computer-based cloning and improvement in codon composition without altering amino acid sequence led to the development of a proposed vaccine. In a nutshell, the findings of this study add to the existing knowledge about the pathogenesis of this infection. The schemed MEV can be a possible prophylactic agent for individuals infected with P. shigelloides. Nevertheless, further authentication is required to guarantee its safeness and immunogenic potential.
Keywords: Codon optimization; Immunoinformatics; Molecular docking; Molecular docking simulation; Plesiomonas shigelloides.
© 2024 Published by Elsevier Ltd.
Conflict of interest statement
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests:Usman Ali Ashfaq reports was provided by Government College University Faisalabad. Usman Ali Ashfaq reports a relationship with Government College University Faisalabad that includes: employment. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


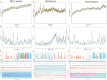
References
-
- Janda J., Family I. Enterobacteriaceae, genus XXVII. Plesiomonas Habs and Schubert 1962. Bergey's manual of systematic bacteriology. 2005;2:740–744.
-
- Ciznar I., et al. Food-Borne Pathogens. Springer; 2006. Plesiomonas shigelloides; pp. 73–80.
-
- Keating J.P. Chronic diarrhea. Pediatr. Rev. 2005;26(1):5–14. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

