Depletion of cap-binding protein eIF4E dysregulates amino acid metabolic gene expression
- PMID: 38848691
- PMCID: PMC11416130
- DOI: 10.1016/j.molcel.2024.05.008
Depletion of cap-binding protein eIF4E dysregulates amino acid metabolic gene expression
Abstract
Protein synthesis is metabolically costly and must be tightly coordinated with changing cellular needs and nutrient availability. The cap-binding protein eIF4E makes the earliest contact between mRNAs and the translation machinery, offering a key regulatory nexus. We acutely depleted this essential protein and found surprisingly modest effects on cell growth and recovery of protein synthesis. Paradoxically, impaired protein biosynthesis upregulated genes involved in the catabolism of aromatic amino acids simultaneously with the induction of the amino acid biosynthetic regulon driven by the integrated stress response factor GCN4. We further identified the translational control of Pho85 cyclin 5 (PCL5), a negative regulator of Gcn4, that provides a consistent protein-to-mRNA ratio under varied translation environments. This regulation depended in part on a uniquely long poly(A) tract in the PCL5 5' UTR and poly(A) binding protein. Collectively, these results highlight how eIF4E connects protein synthesis to metabolic gene regulation, uncovering mechanisms controlling translation during environmental challenges.
Keywords: amino acid metabolism; cap-binding protein; translation.
Copyright © 2024 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests N.T.I. declares equity in Tevard Biosciences and Velia Therapeutics.
Figures






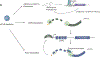
Update of
-
Dysregulation of amino acid metabolism upon rapid depletion of cap-binding protein eIF4E.bioRxiv [Preprint]. 2023 May 12:2023.05.11.540079. doi: 10.1101/2023.05.11.540079. bioRxiv. 2023. Update in: Mol Cell. 2024 Jun 6;84(11):2119-2134.e5. doi: 10.1016/j.molcel.2024.05.008. PMID: 37214807 Free PMC article. Updated. Preprint.
Similar articles
-
Dysregulation of amino acid metabolism upon rapid depletion of cap-binding protein eIF4E.bioRxiv [Preprint]. 2023 May 12:2023.05.11.540079. doi: 10.1101/2023.05.11.540079. bioRxiv. 2023. Update in: Mol Cell. 2024 Jun 6;84(11):2119-2134.e5. doi: 10.1016/j.molcel.2024.05.008. PMID: 37214807 Free PMC article. Updated. Preprint.
-
Rbs1 protein, involved in RNA polymerase III complex assembly in the yeast Saccharomyces cerevisiae, induces a Gcn4 response and forms aggregates when overproduced.Gene. 2022 Jan 30;809:146034. doi: 10.1016/j.gene.2021.146034. Epub 2021 Oct 21. Gene. 2022. PMID: 34688816
-
The 4E-BP Caf20p Mediates Both eIF4E-Dependent and Independent Repression of Translation.PLoS Genet. 2015 May 14;11(5):e1005233. doi: 10.1371/journal.pgen.1005233. eCollection 2015 May. PLoS Genet. 2015. PMID: 25973932 Free PMC article.
-
Translational regulation of GCN4 and the general amino acid control of yeast.Annu Rev Microbiol. 2005;59:407-50. doi: 10.1146/annurev.micro.59.031805.133833. Annu Rev Microbiol. 2005. PMID: 16153175 Review.
-
The Saccharomyces cerevisiae poly (A) binding protein (Pab1): Master regulator of mRNA metabolism and cell physiology.Yeast. 2019 Jan;36(1):23-34. doi: 10.1002/yea.3347. Epub 2018 Oct 17. Yeast. 2019. PMID: 30006991 Review.
Cited by
-
Precise measurement of molecular phenotypes with barcode-based CRISPRi systems.Genome Biol. 2025 May 25;26(1):142. doi: 10.1186/s13059-025-03610-w. Genome Biol. 2025. PMID: 40414878 Free PMC article.
-
eIF4F complex dynamics are important for the activation of the integrated stress response.Mol Cell. 2024 Jun 6;84(11):2135-2151.e7. doi: 10.1016/j.molcel.2024.04.016. Mol Cell. 2024. PMID: 38848692 Free PMC article.
-
Insights into the role of collided ribosomes during the activation of the integrated stress response.Biochem Soc Trans. 2025 Jun 30;53(3):615-626. doi: 10.1042/BST20253034. Biochem Soc Trans. 2025. PMID: 40440025 Free PMC article. Review.
-
Mapping the Genetic Architecture of the Adaptive Integrated Stress Response in S. cerevisiae.bioRxiv [Preprint]. 2024 Dec 22:2024.12.19.629525. doi: 10.1101/2024.12.19.629525. bioRxiv. 2024. PMID: 39763758 Free PMC article. Preprint.
-
CRISPRi with barcoded expression reporters dissects regulatory networks in human cells.bioRxiv [Preprint]. 2024 Sep 6:2024.09.06.611573. doi: 10.1101/2024.09.06.611573. bioRxiv. 2024. PMID: 39282439 Free PMC article. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

