Unveiling novel cell clusters and biomarkers in glioblastoma and its peritumoral microenvironment at the single-cell perspective
- PMID: 38851695
- PMCID: PMC11162569
- DOI: 10.1186/s12967-024-05313-5
Unveiling novel cell clusters and biomarkers in glioblastoma and its peritumoral microenvironment at the single-cell perspective
Abstract
Background: Glioblastoma (GBM) is a highly heterogeneous, recurrent and aggressively invasive primary malignant brain tumor. The heterogeneity of GBM results in poor targeted therapy. Therefore, the aim of this study is to depict the cellular landscape of GBM and its peritumor from a single-cell perspective. Discovering new cell subtypes and biomarkers, and providing a theoretical basis for precision therapy.
Methods: We collected 8 tissue samples from 4 GBM patients to perform 10 × single-cell transcriptome sequencing. Quality control and filtering of data by Seurat package for clustering. Inferring copy number variations to identify malignant cells via the infercnv package. Functional enrichment analysis was performed by GSVA and clusterProfiler packages. STRING database and Cytoscape software were used to construct protein interaction networks. Inferring transcription factors by pySCENIC. Building cell differentiation trajectories via the monocle package. To infer intercellular communication networks by CellPhoneDB software.
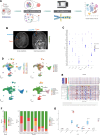
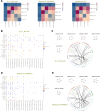
Results: We observed that the tumor microenvironment (TME) varies among different locations and different GBM patients. We identified a proliferative cluster of oligodendrocytes with high expression of mitochondrial genes. We also identified two clusters of myeloid cells, one primarily located in the peritumor exhibiting an M1 phenotype with elevated TNFAIP8L3 expression, and another in the tumor and peritumor showing a proliferative tendency towards an M2 phenotype with increased DTL expression. We identified XIST, KCNH7, SYT1 and DIAPH3 as potential factors associated with the proliferation of malignant cells in GBM.
Conclusions: These biomarkers and cell clusters we discovered may serve as targets for treatment. Targeted drugs developed against these biomarkers and cell clusters may enhance treatment efficacy, optimize immune therapy strategies, and improve the response rates of GBM patients to immunotherapy. Our findings provide a theoretical basis for the development of individualized treatment and precision medicine for GBM, which may be used to improve the survival of GBM patients.
Keywords: Biomarkers; Glioblastoma; Single-cell sequencing; Tumor heterogeneity; Tumor microenvironment.
© 2024. The Author(s).
Conflict of interest statement
The authors have declared that no competing interest exists.
Figures






Similar articles
-
Exploring the prognostic value of BRMS1 + microglia based on single-cell anoikis regulator patterns in the immunologic microenvironment of GBM.J Neurooncol. 2024 Oct;170(1):101-117. doi: 10.1007/s11060-024-04781-5. Epub 2024 Aug 15. J Neurooncol. 2024. PMID: 39143438 Free PMC article.
-
Unveiling the NEFH+ malignant cell subtype: Insights from single-cell RNA sequencing in prostate cancer progression and tumor microenvironment interactions.Front Immunol. 2024 Dec 20;15:1517679. doi: 10.3389/fimmu.2024.1517679. eCollection 2024. Front Immunol. 2024. PMID: 39759507 Free PMC article.
-
Machine learning and multi-omics analysis reveal key regulators of proneural-mesenchymal transition in glioblastoma.Sci Rep. 2025 Jun 5;15(1):19731. doi: 10.1038/s41598-025-04862-z. Sci Rep. 2025. PMID: 40473799 Free PMC article.
-
"Zooming in" on Glioblastoma: Understanding Tumor Heterogeneity and its Clinical Implications in the Era of Single-Cell Ribonucleic Acid Sequencing.Neurosurgery. 2021 Feb 16;88(3):477-486. doi: 10.1093/neuros/nyaa305. Neurosurgery. 2021. PMID: 32674143 Review.
-
Comprehensive understanding of glioblastoma molecular phenotypes: classification, characteristics, and transition.Cancer Biol Med. 2024 May 6;21(5):363-81. doi: 10.20892/j.issn.2095-3941.2023.0510. Cancer Biol Med. 2024. PMID: 38712813 Free PMC article. Review.
Cited by
-
Decoding the Tumor Microenvironment of Myoepithelial Cells in Triple-Negative Breast Cancer Through Single-Cell and Transcriptomic Sequencing and Establishing a Prognostic Model Based on Key Myoepithelial Cell Genes.Int J Genomics. 2025 May 6;2025:6454413. doi: 10.1155/ijog/6454413. eCollection 2025. Int J Genomics. 2025. PMID: 40365116 Free PMC article.
-
Multi-Omics Integration: Predicting Progression and Optimizing Clinical Treatment of Hepatocellular Carcinoma Through Malignant-Cell-Related Genes.Int J Mol Sci. 2025 Jun 26;26(13):6135. doi: 10.3390/ijms26136135. Int J Mol Sci. 2025. PMID: 40649911 Free PMC article.
-
Vagus nerve stimulation: Novel concept for the treatment of glioblastoma and solid cancers by cytokine (interleukin-6) reduction, attenuating the SASP, enhancing tumor immunity.Brain Behav Immun Health. 2024 Sep 17;42:100859. doi: 10.1016/j.bbih.2024.100859. eCollection 2024 Dec. Brain Behav Immun Health. 2024. PMID: 39512605 Free PMC article. Review.
-
TNFAIP8L3 regulation of the TGF-β signaling pathway affects the proportion of macrophages during tumor antigen presentation and affects the prognosis of ovarian cancer.Front Mol Biosci. 2025 Apr 24;12:1566363. doi: 10.3389/fmolb.2025.1566363. eCollection 2025. Front Mol Biosci. 2025. PMID: 40343258 Free PMC article.
-
Identification and validation of tricarboxylic acid cycle-related diagnostic biomarkers for diabetic nephropathy via weighted gene co-expression network analysis and single-cell transcriptome analysis.Acta Diabetol. 2025 Jul 31. doi: 10.1007/s00592-025-02557-5. Online ahead of print. Acta Diabetol. 2025. PMID: 40742465
References
-
- Fernandes C, Costa A, Osório L, Lago RC, Linhares P, Carvalho B, et al. Current standards of care in glioblastoma therapy. In: De Vleeschouwer S, et al., editors. Glioblastoma. Brisbane (AU): Codon Publications Copyright: The Authors; 2017. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

