Effects of highly active antiretroviral therapy initiation on epigenomic DNA methylation in persons living with HIV
- PMID: 38855142
- PMCID: PMC11157437
- DOI: 10.3389/fbinf.2024.1357889
Effects of highly active antiretroviral therapy initiation on epigenomic DNA methylation in persons living with HIV
Abstract
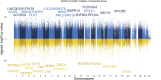
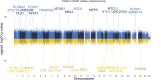
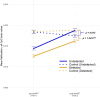
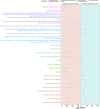
Introduction: Highly active antiretroviral therapy (HAART) helps improve some measures of accelerated epigenetic aging in persons living with HIV (PLWH), but its overall impact on the epigenome is not fully understood. Methods: In this study, we analyzed the DNA methylation profiles of PLWH (n = 187) shortly before and approximately 2-3 years after they started HAART, as well as matched seronegative (SN) controls (n = 187), taken at two time intervals. Our aim was to identify specific CpGs and biologic pathways associated with HIV infection and initiation of HAART. Additionally, we attempted to identify epigenetic changes associated with HAART initiation that were independent of HIV-associated changes, using matched HIV seronegative (SN) controls (matched on age, hepatitis C status, and interval between visits) to identify CpGs that did not differ between PLWH and SN pre-HAART but were significantly associated with HAART initiation while being unrelated to HIV viral load. Epigenome-wide association studies (EWAS) on >850,000 CpG sites were performed using pre- and post-HAART samples from PLWH. The results were then annotated using the Genomic Regions Enrichment of Annotations Tool (GREAT). Results: When only pre- and post-HAART visits in PLWH were compared, gene ontologies related to immune function and diseases related to immune function were significant, though with less significance for PLWH with detectable HIV viral loads (>50 copies/mL) at the post-HAART visit. To specifically elucidate the effects of HAART separately from HIV-induced methylation changes, we performed EWAS of HAART while also controlling for HIV viral load, and found gene ontologies associated with transplant rejection, transplant-related diseases, and other immunologic signatures. Additionally, we performed a more focused analysis that examined CpGs reaching genome-wide significance (p < 1 × 10-7) from the viral load-controlled EWAS that did not differ between all PLWH and matched SN controls pre-HAART. These CpGs were found to be near genes that play a role in retroviral drug metabolism, diffuse large B cell lymphoma proliferation, and gastric cancer metastasis. Discussion: Overall, this study provides insight into potential biological functions associated with DNA methylation changes induced by HAART initiation in persons living with HIV.
Keywords: DNA methylation; EWAS; HIV; antiretroviral therapy; bioinformatics; epigenetics.
Copyright © 2024 Zhang, Sehl, Shih, Breen, Li, Lu, Bream, Duggal, Martinson, Wolinsky, Martinez-Maza, Ramirez, Horvath and Jamieson.
Conflict of interest statement
SH is a founder of the non-profit Epigenetic Clock Development Foundation which plans to license several patents from his employer UC Regents. These patents list SH as inventor. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. CR declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures




Similar articles
-
Decreased but persistent epigenetic age acceleration is associated with changes in T-cell subsets after initiation of highly active antiretroviral therapy in persons living with HIV.Front Bioinform. 2024 May 24;4:1356509. doi: 10.3389/fbinf.2024.1356509. eCollection 2024. Front Bioinform. 2024. PMID: 38855141 Free PMC article.
-
Increased Rate of Epigenetic Aging in Men Living With HIV Prior to Treatment.Front Genet. 2022 Feb 28;12:796547. doi: 10.3389/fgene.2021.796547. eCollection 2021. Front Genet. 2022. PMID: 35295196 Free PMC article.
-
Kidney and liver organ transplantation in persons with human immunodeficiency virus: An Evidence-Based Analysis.Ont Health Technol Assess Ser. 2010;10(4):1-56. Epub 2010 Mar 1. Ont Health Technol Assess Ser. 2010. PMID: 23074407 Free PMC article.
-
High-sensitivity C-reactive protein among people living with HIV on highly active antiretroviral therapy: a systemic review and meta-analysis.BMC Infect Dis. 2024 Feb 2;24(1):160. doi: 10.1186/s12879-024-09050-4. BMC Infect Dis. 2024. PMID: 38308222 Free PMC article.
-
Molecular biological assessment methods and understanding the course of the HIV infection.APMIS Suppl. 2003;(114):1-37. APMIS Suppl. 2003. PMID: 14626050 Review.
Cited by
-
Latency Reversing Agents and the Road to an HIV Cure.Pathogens. 2025 Feb 27;14(3):232. doi: 10.3390/pathogens14030232. Pathogens. 2025. PMID: 40137717 Free PMC article. Review.
-
PON-1 and PON-2 Polymorphisms and PON-1 Paraoxonase Activity in People Living with HIV-1.Antioxidants (Basel). 2025 Feb 12;14(2):209. doi: 10.3390/antiox14020209. Antioxidants (Basel). 2025. PMID: 40002395 Free PMC article.
References
-
- Aryee M. J., Jaffe A. E., Corrada-Bravo H., Ladd-Acosta C., Feinberg A. P., Hansen K. D., et al. (2014). Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30 (10), 1363–1369. 10.1093/bioinformatics/btu049 - DOI - PMC - PubMed
-
- Bhagat P., Kaur Sachdeva R., Sharma P., Updesh Singh Sachdeva M., Chhabra S., Sharma A., et al. (2015). Effect of antiretroviral therapy on hemoglobin A2 values can have implications in antenatal beta-thalassemia screening programs. Infect. Dis. 48 (2), 122–126. 10.3109/23744235.2015.1089592 - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

