Histone acetyltransferase Kat2a regulates ferroptosis via enhancing Tfrc and Hmox1 expression in diabetic cardiomyopathy
- PMID: 38858351
- PMCID: PMC11164963
- DOI: 10.1038/s41419-024-06771-x
Histone acetyltransferase Kat2a regulates ferroptosis via enhancing Tfrc and Hmox1 expression in diabetic cardiomyopathy
Abstract
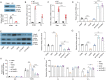
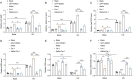
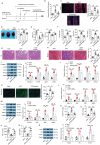
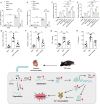
Diabetic cardiomyopathy (DCM) is a prevalent myocardial microvascular complication of the myocardium with a complex pathogenesis. Investigating the pathogenesis of DCM can significantly contribute to enhancing its prevention and treatment strategies. Our study revealed an upregulation of lysine acetyltransferase 2 A (Kat2a) expression in DCM, accompanied by a decrease in N6-methyladenosine (m6A) modified Kat2a mRNA levels. Our study revealed an upregulation of lysine acetyltransferase 2 A (Kat2a) expression in DCM, accompanied by a decrease in N6-methyladenosine (m6A) modified Kat2a mRNA levels. Functionally, inhibition of Kat2a effectively ameliorated high glucose-induced cardiomyocyte injury both in vitro and in vivo by suppressing ferroptosis. Mechanistically, Demethylase alkB homolog 5 (Alkbh5) was found to reduce m6A methylation levels on Kat2a mRNA, leading to its upregulation. YTH domain family 2 (Ythdf2) played a crucial role as an m6A reader protein mediating the degradation of Kat2a mRNA. Furthermore, Kat2a promoted ferroptosis by increasing Tfrc and Hmox1 expression via enhancing the enrichment of H3K27ac and H3K9ac on their promoter regions. In conclusion, our findings unveil a novel role for the Kat2a-ferroptosis axis in DCM pathogenesis, providing valuable insights for potential clinical interventions.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
miR-340-3p-modified bone marrow mesenchymal stem cell-derived exosomes inhibit ferroptosis through METTL3-mediated m6A modification of HMOX1 to promote recovery of injured rat uterus.Stem Cell Res Ther. 2024 Jul 29;15(1):224. doi: 10.1186/s13287-024-03846-6. Stem Cell Res Ther. 2024. PMID: 39075530 Free PMC article.
-
m6A demethylase ALKBH5 reduces ferroptosis in diabetic retinopathy through the m6A-YTHDF1-ACSL4 axis.Diabet Med. 2025 Aug;42(8):e70033. doi: 10.1111/dme.70033. Epub 2025 Apr 10. Diabet Med. 2025. PMID: 40210448
-
N6-methyladenosine modification of SPOP relieves ferroptosis and diabetic cardiomyopathy by enhancing ubiquitination of VDAC3.Free Radic Biol Med. 2025 Jan;226:216-229. doi: 10.1016/j.freeradbiomed.2024.11.025. Epub 2024 Nov 15. Free Radic Biol Med. 2025. PMID: 39549880
-
Potential Significance of Targeting Ferroptosis for Intervention of Diabetic Cardiomyopathy.J Diabetes. 2025 Jun;17(6):e70116. doi: 10.1111/1753-0407.70116. J Diabetes. 2025. PMID: 40536196 Free PMC article. Review.
-
The Interplay of N6-Methyladenosine and Ferroptosis in Cancer: A Promising Therapeutic Avenue.Cancer Genet. 2025 Sep;296-297:15-24. doi: 10.1016/j.cancergen.2025.05.007. Epub 2025 Jun 1. Cancer Genet. 2025. PMID: 40482423 Review.
Cited by
-
Ferroptosis in diabetic cardiomyopathy: from its mechanisms to therapeutic strategies.Front Endocrinol (Lausanne). 2024 Nov 11;15:1421838. doi: 10.3389/fendo.2024.1421838. eCollection 2024. Front Endocrinol (Lausanne). 2024. PMID: 39588340 Free PMC article. Review.
-
Ferroptosis: mechanism and role in diabetes-related cardiovascular diseases.Cardiovasc Diabetol. 2025 Feb 7;24(1):60. doi: 10.1186/s12933-025-02614-x. Cardiovasc Diabetol. 2025. PMID: 39920799 Free PMC article. Review.
-
Integrative Mendelian randomization and multi-omics analysis identifies anti-allergic drug targets associated with cardiovascular disease risk.Sci Rep. 2025 Aug 21;15(1):30783. doi: 10.1038/s41598-025-15331-y. Sci Rep. 2025. PMID: 40841572 Free PMC article.
-
The m6A modification of SOX18 leads to increased PTX3 and cardiomyocyte pyroptosis in sepsis-induced cardiomyopathy.Theranostics. 2025 Feb 24;15(8):3532-3550. doi: 10.7150/thno.103809. eCollection 2025. Theranostics. 2025. PMID: 40093897 Free PMC article.
-
Recent advances associated with cardiometabolic remodeling in diabetes-induced heart failure.Am J Physiol Heart Circ Physiol. 2024 Dec 1;327(6):H1327-H1342. doi: 10.1152/ajpheart.00539.2024. Epub 2024 Oct 25. Am J Physiol Heart Circ Physiol. 2024. PMID: 39453429 Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

