PR-SET7 epigenetically restrains uterine interferon response and cell death governing proper postnatal stromal development
- PMID: 38858353
- PMCID: PMC11164956
- DOI: 10.1038/s41467-024-49342-6
PR-SET7 epigenetically restrains uterine interferon response and cell death governing proper postnatal stromal development
Abstract
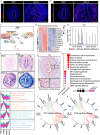
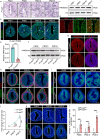
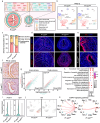
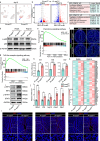
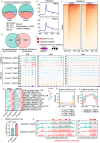
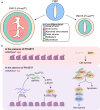
The differentiation of the stroma is a hallmark event during postnatal uterine development. However, the spatiotemporal changes that occur during this process and the underlying regulatory mechanisms remain elusive. Here, we comprehensively delineated the dynamic development of the neonatal uterus at single-cell resolution and characterized two distinct stromal subpopulations, inner and outer stroma. Furthermore, single-cell RNA sequencing revealed that uterine ablation of Pr-set7, the sole methyltransferase catalyzing H4K20me1, led to a reduced proportion of the inner stroma due to massive cell death, thus impeding uterine development. By combining RNA sequencing and epigenetic profiling of H4K20me1, we demonstrated that PR-SET7-H4K20me1 either directly repressed the transcription of interferon stimulated genes or indirectly restricted the interferon response via silencing endogenous retroviruses. Declined H4K20me1 level caused viral mimicry responses and ZBP1-mediated apoptosis and necroptosis in stromal cells. Collectively, our study provides insight into the epigenetic machinery governing postnatal uterine stromal development mediated by PR-SET7.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
PR-Set7 deficiency limits uterine epithelial population growth hampering postnatal gland formation in mice.Cell Death Differ. 2017 Dec;24(12):2013-2021. doi: 10.1038/cdd.2017.120. Epub 2017 Jul 21. Cell Death Differ. 2017. PMID: 28731465 Free PMC article.
-
Trophoblast PR-SET7 dysfunction induces viral mimicry response and necroptosis associated with recurrent miscarriage.Proc Natl Acad Sci U S A. 2023 Jun 20;120(25):e2216206120. doi: 10.1073/pnas.2216206120. Epub 2023 Jun 12. Proc Natl Acad Sci U S A. 2023. PMID: 37307441 Free PMC article.
-
The PR-Set7 binding domain of Riz1 is required for the H4K20me1-H3K9me1 trans-tail 'histone code' and Riz1 tumor suppressor function.Nucleic Acids Res. 2014 Apr;42(6):3580-9. doi: 10.1093/nar/gkt1377. Epub 2014 Jan 13. Nucleic Acids Res. 2014. PMID: 24423864 Free PMC article.
-
[The biological functions of lysine methyltransferase PR-SET7].Yi Chuan. 2013 Mar;35(3):241-54. doi: 10.3724/sp.j.1005.2013.00241. Yi Chuan. 2013. PMID: 23575530 Review. Chinese.
-
PR-Set7 and H4K20me1: at the crossroads of genome integrity, cell cycle, chromosome condensation, and transcription.Genes Dev. 2012 Feb 15;26(4):325-37. doi: 10.1101/gad.177444.111. Genes Dev. 2012. PMID: 22345514 Free PMC article. Review.
Cited by
-
Histone H4 lysine 20 monomethylation is not a mark of transcriptional silencers.bioRxiv [Preprint]. 2025 Jan 13:2025.01.09.632211. doi: 10.1101/2025.01.09.632211. bioRxiv. 2025. PMID: 39868205 Free PMC article. Preprint.
-
Decoding Müllerian Duct Epithelial Regionalization.Mol Reprod Dev. 2025 Feb;92(2):e70018. doi: 10.1002/mrd.70018. Mol Reprod Dev. 2025. PMID: 39994938 Free PMC article. Review.
-
Uterine organoids reveal insights into epithelial specification and plasticity in development and disease.Proc Natl Acad Sci U S A. 2025 Feb 4;122(5):e2422694122. doi: 10.1073/pnas.2422694122. Epub 2025 Jan 30. Proc Natl Acad Sci U S A. 2025. PMID: 39883834 Free PMC article.
-
WT1 directs normal progesterone receptor-chromatin binding essential for uterine receptivity at peri-implantation.Proc Natl Acad Sci U S A. 2025 Jul 15;122(28):e2504361122. doi: 10.1073/pnas.2504361122. Epub 2025 Jul 8. Proc Natl Acad Sci U S A. 2025. PMID: 40627402
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

