OptoLacI: optogenetically engineered lactose operon repressor LacI responsive to light instead of IPTG
- PMID: 38860425
- PMCID: PMC11260447
- DOI: 10.1093/nar/gkae479
OptoLacI: optogenetically engineered lactose operon repressor LacI responsive to light instead of IPTG
Abstract
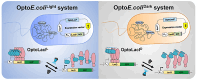
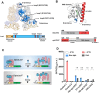
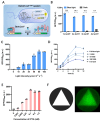
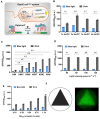
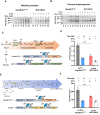
Optogenetics' advancement has made light induction attractive for controlling biological processes due to its advantages of fine-tunability, reversibility, and low toxicity. The lactose operon induction system, commonly used in Escherichia coli, relies on the binding of lactose or isopropyl β-d-1-thiogalactopyranoside (IPTG) to the lactose repressor protein LacI, playing a pivotal role in controlling the lactose operon. Here, we harnessed the light-responsive light-oxygen-voltage 2 (LOV2) domain from Avena sativa phototropin 1 as a tool for light control and engineered LacI into two light-responsive variants, OptoLacIL and OptoLacID. These variants exhibit direct responsiveness to light and darkness, respectively, eliminating the need for IPTG. Building upon OptoLacI, we constructed two light-controlled E. coli gene expression systems, OptoE.coliLight system and OptoE.coliDark system. These systems enable bifunctional gene expression regulation in E. coli through light manipulation and show superior controllability compared to IPTG-induced systems. We applied the OptoE.coliDark system to protein production and metabolic flux control. Protein production levels are comparable to those induced by IPTG. Notably, the titers of dark-induced production of 1,3-propanediol (1,3-PDO) and ergothioneine exceeded 110% and 60% of those induced by IPTG, respectively. The development of OptoLacI will contribute to the advancement of the field of optogenetic protein engineering, holding substantial potential applications across various fields.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
Similar articles
-
Optogenetic control of the lac operon for bacterial chemical and protein production.Nat Chem Biol. 2021 Jan;17(1):71-79. doi: 10.1038/s41589-020-0639-1. Epub 2020 Sep 7. Nat Chem Biol. 2021. PMID: 32895498
-
Establishment of a low-dosage-IPTG inducible expression system construction method in Escherichia coli.J Basic Microbiol. 2018 Sep;58(9):806-810. doi: 10.1002/jobm.201800160. Epub 2018 Jul 2. J Basic Microbiol. 2018. PMID: 29962051
-
A single mutation in the core domain of the lac repressor reduces leakiness.Microb Cell Fact. 2013 Jul 8;12:67. doi: 10.1186/1475-2859-12-67. Microb Cell Fact. 2013. PMID: 23834731 Free PMC article.
-
Lac repressor-operator complex.Curr Opin Struct Biol. 1997 Feb;7(1):76-85. doi: 10.1016/s0959-440x(97)80010-3. Curr Opin Struct Biol. 1997. PMID: 9032054 Review.
-
Review: optimizing inducer and culture conditions for expression of foreign proteins under the control of the lac promoter.J Ind Microbiol. 1996 Mar;16(3):145-54. doi: 10.1007/BF01569997. J Ind Microbiol. 1996. PMID: 8652113 Review.
Cited by
-
Induction of bacterial expression at the mRNA level by light.Nucleic Acids Res. 2024 Sep 9;52(16):10017-10028. doi: 10.1093/nar/gkae678. Nucleic Acids Res. 2024. PMID: 39126322 Free PMC article.
-
Optogenetic control of Corynebacterium glutamicum gene expression.Nucleic Acids Res. 2024 Dec 11;52(22):14260-14276. doi: 10.1093/nar/gkae1149. Nucleic Acids Res. 2024. PMID: 39607706 Free PMC article.
References
-
- Lewis M., Chang G., Horton N.C., Kercher M.A., Pace H.C., Schumacher M.A., Brennan R.G., Lu P. Crystal structure of the lactose operon repressor and its complexes with DNA and inducer. Science. 1996; 271:1247–1254. - PubMed
-
- Jacob F., Monod J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mole. Biol. 1961; 3:318–356. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

