MYG1 drives glycolysis and colorectal cancer development through nuclear-mitochondrial collaboration
- PMID: 38862489
- PMCID: PMC11167044
- DOI: 10.1038/s41467-024-49221-0
MYG1 drives glycolysis and colorectal cancer development through nuclear-mitochondrial collaboration
Abstract
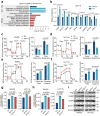
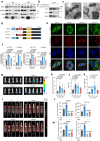
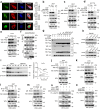
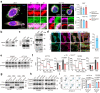
Metabolic remodeling is a strategy for tumor survival under stress. However, the molecular mechanisms during the metabolic remodeling of colorectal cancer (CRC) remain unclear. Melanocyte proliferating gene 1 (MYG1) is a 3'-5' RNA exonuclease and plays a key role in mitochondrial functions. Here, we uncover that MYG1 expression is upregulated in CRC progression and highly expressed MYG1 promotes glycolysis and CRC progression independent of its exonuclease activity. Mechanistically, nuclear MYG1 recruits HSP90/GSK3β complex to promote PKM2 phosphorylation, increasing its stability. PKM2 transcriptionally activates MYC and promotes MYC-medicated glycolysis. Conversely, c-Myc also transcriptionally upregulates MYG1, driving the progression of CRC. Meanwhile, mitochondrial MYG1 on the one hand inhibits oxidative phosphorylation (OXPHOS), and on the other hand blocks the release of Cyt c from mitochondria and inhibits cell apoptosis. Clinically, patients with KRAS mutation show high expression of MYG1, indicating a high level of glycolysis and a poor prognosis. Targeting MYG1 may disturb metabolic balance of CRC and serve as a potential target for the diagnosis and treatment of CRC.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- International Agency for Research on Cancer. Globocan 2020: Cancer Fact Sheets—Colorectal Cancer. IARC [Internet]. Available from: http://gco.iarc.fr/today/data/factsheets/cancers/10_8_9-Colorectum-fact-....
-
- Hanahan D. Hallmarks of cancer: new dimensions. Cancer Discov. 2022;12:31–46. doi: 10.1158/2159-8290.CD-21-1059. - DOI - PubMed
MeSH terms
Substances
Grants and funding
- 81272763/National Natural Science Foundation of China (National Science Foundation of China)
- 81672466/National Natural Science Foundation of China (National Science Foundation of China)
- 81972334/National Natural Science Foundation of China (National Science Foundation of China)
- 82173297/National Natural Science Foundation of China (National Science Foundation of China)
- 82373066/National Natural Science Foundation of China (National Science Foundation of China)
- 82273351/National Natural Science Foundation of China (National Science Foundation of China)
- 2017A030313550/Natural Science Foundation of Guangdong Province (Guangdong Natural Science Foundation)
- 2019A1515011205/Natural Science Foundation of Guangdong Province (Guangdong Natural Science Foundation)
- 2023A1515010329/Natural Science Foundation of Guangdong Province (Guangdong Natural Science Foundation)
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

