Salinity-responsive histone PTMs identified in the gills and gonads of Mozambique tilapia (Oreochromis mossambicus)
- PMID: 38862901
- PMCID: PMC11167857
- DOI: 10.1186/s12864-024-10471-3
Salinity-responsive histone PTMs identified in the gills and gonads of Mozambique tilapia (Oreochromis mossambicus)
Abstract
Background: Histone post-translational modifications (PTMs) are epigenetic marks that can be induced by environmental stress and elicit heritable patterns of gene expression. To investigate this process in an ecological context, we characterized the influence of salinity stress on histone PTMs within the gills, kidney, and testes of Mozambique tilapia (Oreochromis mossambicus). A total of 221 histone PTMs were quantified in each tissue sample and compared between freshwater-adapted fish exposed to salinity treatments that varied in intensity and duration.
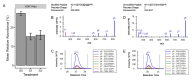
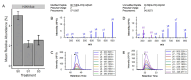
Results: Four salinity-responsive histone PTMs were identified in this study. When freshwater-adapted fish were exposed to seawater for two hours, the relative abundance of H1K16ub significantly increased in the gills. Long-term salinity stress elicited changes in both the gills and testes. When freshwater-adapted fish were exposed to a pulse of severe salinity stress, where salinity gradually increased from freshwater to a maximum of 82.5 g/kg, the relative abundance of H1S1ac significantly decreased in the gills. Under the same conditions, the relative abundance of both H3K14ac and H3K18ub decreased significantly in the testes of Mozambique tilapia.
Conclusions: This study demonstrates that salinity stress can alter histone PTMs in the gills and gonads of Mozambique tilapia, which, respectively, signify a potential for histone PTMs to be involved in salinity acclimation and adaptation in euryhaline fishes. These results thereby add to a growing body of evidence that epigenetic mechanisms may be involved in such processes.
Keywords: Epigenetics; Euryhaline fish; Mass spectrometry; Salinity stress; Stress-induced evolution.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
Environmental conditions elicit a slow but enduring response of histone post-translational modifications in Mozambique tilapia.Environ Epigenet. 2024 Sep 2;10(1):dvae013. doi: 10.1093/eep/dvae013. eCollection 2024. Environ Epigenet. 2024. PMID: 39372708 Free PMC article.
-
A Strategy to Characterize the Global Landscape of Histone Post-Translational Modifications Within Tissues of Nonmodel Organisms.J Proteome Res. 2024 Aug 2;23(8):2780-2794. doi: 10.1021/acs.jproteome.3c00246. Epub 2023 Aug 25. J Proteome Res. 2024. PMID: 37624673 Free PMC article.
-
Differential transcriptomic analyses revealed genes and signaling pathways involved in iono-osmoregulation and cellular remodeling in the gills of euryhaline Mozambique tilapia, Oreochromis mossambicus.BMC Genomics. 2014 Oct 23;15(1):921. doi: 10.1186/1471-2164-15-921. BMC Genomics. 2014. PMID: 25342237 Free PMC article.
-
Characterization and expression of Na+/K+-ATPase in gills and kidneys of the Teleost fish Oreochromis mossambicus, Oreochromis urolepis hornorum and their hybrids in response to salinity challenge.Comp Biochem Physiol A Mol Integr Physiol. 2018 Oct;224:1-10. doi: 10.1016/j.cbpa.2018.05.017. Epub 2018 May 28. Comp Biochem Physiol A Mol Integr Physiol. 2018. PMID: 29852253
-
Endocrine and osmoregulatory responses to tidally-changing salinities in fishes.Gen Comp Endocrinol. 2022 Sep 15;326:114071. doi: 10.1016/j.ygcen.2022.114071. Epub 2022 Jun 11. Gen Comp Endocrinol. 2022. PMID: 35697315 Review.
Cited by
-
Environmental conditions elicit a slow but enduring response of histone post-translational modifications in Mozambique tilapia.Environ Epigenet. 2024 Sep 2;10(1):dvae013. doi: 10.1093/eep/dvae013. eCollection 2024. Environ Epigenet. 2024. PMID: 39372708 Free PMC article.
-
Adaptation responses to salt stress in the gut of Poecilia reticulata.Anim Cells Syst (Seoul). 2025 Jan 18;29(1):84-99. doi: 10.1080/19768354.2025.2451413. eCollection 2025. Anim Cells Syst (Seoul). 2025. PMID: 39839657 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

