ERRFI1 exacerbates hepatic ischemia reperfusion injury by promoting hepatocyte apoptosis and ferroptosis in a GRB2-dependent manner
- PMID: 38862918
- PMCID: PMC11167874
- DOI: 10.1186/s10020-024-00837-4
ERRFI1 exacerbates hepatic ischemia reperfusion injury by promoting hepatocyte apoptosis and ferroptosis in a GRB2-dependent manner
Abstract
Background: Programmed cell death is an important mechanism for the development of hepatic ischemia and reperfusion (IR) injury, and multiple novel forms of programmed cell death are involved in the pathological process of hepatic IR. ERRFI1 is involved in the regulation of cell apoptosis in myocardial IR. However, the function of ERRFI1 in hepatic IR injury and its modulation of programmed cell death remain largely unknown.
Methods: Here, we performed functional and molecular mechanism studies in hepatocyte-specific knockout mice and ERRFI1-silenced hepatocytes to investigate the significance of ERRFI1 in hepatic IR injury. The histological severity of livers, enzyme activities, hepatocyte apoptosis and ferroptosis were determined.
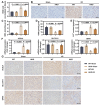
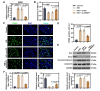
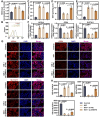
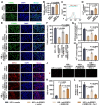
Results: ERRFI1 expression increased in liver tissues from mice with IR injury and hepatocytes under oxygen-glucose deprivation/reoxygenation (OGD/R) conditions. Hepatocyte-specific ERRFI1 knockout alleviated IR-induced liver injury in mice by reducing cell apoptosis and ferroptosis. ERRFI1 knockdown reduced apoptotic and ferroptotic hepatocytes induced by OGD/R. Mechanistically, ERRFI1 interacted with GRB2 to maintain its stability by hindering its proteasomal degradation. Overexpression of GRB2 abrogated the effects of ERRFI1 silencing on hepatocyte apoptosis and ferroptosis.
Conclusions: Our results revealed that the ERRFI1-GRB2 interaction and GRB2 stability are essential for ERRFI1-regulated hepatic IR injury, indicating that inhibition of ERRFI1 or blockade of the ERRFI1-GRB2 interaction may be potential therapeutic strategies in response to hepatic IR injury.
Keywords: Apoptosis; ERRFI1; Ferroptosis; GRB2; Hepatic ischemia and reperfusion injury.
© 2024. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
Hepatocyte-specific TMEM16A deficiency alleviates hepatic ischemia/reperfusion injury via suppressing GPX4-mediated ferroptosis.Cell Death Dis. 2022 Dec 26;13(12):1072. doi: 10.1038/s41419-022-05518-w. Cell Death Dis. 2022. PMID: 36572666 Free PMC article.
-
TRIM21 knockdown alleviates hemorrhage induced hepatic ischemia reperfusion injury by suppressing ferroptosis-induced NETs.Sci Rep. 2025 May 14;15(1):16731. doi: 10.1038/s41598-025-99182-7. Sci Rep. 2025. PMID: 40369118 Free PMC article.
-
Indoleamine 2, 3-dioxygenase 1 activation in macrophage exacerbates hepatic ischemia-reperfusion injury by triggering hepatocyte ferroptosis.Int Immunopharmacol. 2024 Mar 30;130:111692. doi: 10.1016/j.intimp.2024.111692. Epub 2024 Feb 20. Int Immunopharmacol. 2024. PMID: 38382261
-
Ferroptosis: a new target for hepatic ischemia-reperfusion injury?Free Radic Res. 2024 May-Jun;58(6-7):396-416. doi: 10.1080/10715762.2024.2386075. Epub 2024 Jul 31. Free Radic Res. 2024. PMID: 39068663 Review.
-
Programmed cell death, from liver Ischemia-Reperfusion injury perspective: An overview.Heliyon. 2024 Jun 17;10(13):e32480. doi: 10.1016/j.heliyon.2024.e32480. eCollection 2024 Jul 15. Heliyon. 2024. PMID: 39040334 Free PMC article. Review.
Cited by
-
Identification of Grb2 protein as a potential mediator of macrophage activation in acute pancreatitis based on bioinformatics and experimental verification.Front Immunol. 2025 May 26;16:1575880. doi: 10.3389/fimmu.2025.1575880. eCollection 2025. Front Immunol. 2025. PMID: 40491924 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

