Decoding frontotemporal and cell-type-specific vulnerabilities to neuropsychiatric disorders and psychoactive drugs
- PMID: 38864245
- PMCID: PMC11285532
- DOI: 10.1098/rsob.240063
Decoding frontotemporal and cell-type-specific vulnerabilities to neuropsychiatric disorders and psychoactive drugs
Abstract
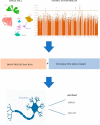
Frontotemporal lobe abnormalities are linked to neuropsychiatric disorders and cognition, but the role of cellular heterogeneity between temporal lobe (TL) and frontal lobe (FL) in the vulnerability to genetic risk factors remains to be elucidated. We integrated single-nucleus transcriptome analysis in 'fresh' human FL and TL with genetic susceptibility, gene dysregulation in neuropsychiatric disease and psychoactive drug response data. We show how intrinsic differences between TL and FL contribute to the vulnerability of specific cell types to both genetic risk factors and psychoactive drugs. Neuronal populations, specifically PVALB neurons, were most highly vulnerable to genetic risk factors for psychiatric disease. These psychiatric disease-associated genes were mostly upregulated in the TL, and dysregulated in the brain of patients with obsessive-compulsive disorder, bipolar disorder and schizophrenia. Among these genes, GRIN2A and SLC12A5, implicated in schizophrenia and bipolar disorder, were significantly upregulated in TL PVALB neurons and in psychiatric disease patients' brain. PVALB neurons from the TL were twofold more vulnerable to psychoactive drugs than to genetic risk factors, showing the influence and specificity of frontotemporal lobe differences on cell vulnerabilities. These studies provide a cell type resolved map of the impact of brain regional differences on cell type vulnerabilities in neuropsychiatric disorders.
Keywords: frontal lobe; psychiatric disorder; psychoactive drugs; single-cell RNA sequencing; temporal lobe.
Conflict of interest statement
We declare we have no competing interests.
Figures





Similar articles
-
Transcriptomic pathology of neocortical microcircuit cell types across psychiatric disorders.Mol Psychiatry. 2025 Mar;30(3):1057-1068. doi: 10.1038/s41380-024-02707-1. Epub 2024 Sep 5. Mol Psychiatry. 2025. PMID: 39237723
-
Choline Compounds of the Frontal Lobe and Temporal Glutamatergic System in Bipolar and Schizophrenia Proton Magnetic Resonance Spectroscopy Study.Dis Markers. 2018 Nov 25;2018:3654894. doi: 10.1155/2018/3654894. eCollection 2018. Dis Markers. 2018. PMID: 30595760 Free PMC article.
-
N-acetylaspartate and N-Acetylaspartylglutamate deficits in superior temporal cortex in schizophrenia and bipolar disorder: a postmortem study.Biol Psychiatry. 2003 Jun 15;53(12):1138-41. doi: 10.1016/s0006-3223(02)01742-0. Biol Psychiatry. 2003. PMID: 12814865
-
Postmortem human brain genomics in neuropsychiatric disorders--how far can we go?Curr Opin Neurobiol. 2016 Feb;36:107-11. doi: 10.1016/j.conb.2015.11.002. Epub 2015 Dec 10. Curr Opin Neurobiol. 2016. PMID: 26685806 Free PMC article. Review.
-
Epigenetics of psychoactive drugs.J Pharm Pharmacol. 2012 Oct;64(10):1349-58. doi: 10.1111/j.2042-7158.2012.01475.x. Epub 2012 Apr 8. J Pharm Pharmacol. 2012. PMID: 22943166 Review.
Cited by
-
Brain connectivity and transcriptional changes induced by rTMS in first-episode major depressive disorder.Transl Psychiatry. 2025 Apr 24;15(1):159. doi: 10.1038/s41398-025-03376-6. Transl Psychiatry. 2025. PMID: 40274783 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

