Super-enhancer-driven IRF2BP2 enhances ALK activity and promotes neuroblastoma cell proliferation
- PMID: 38864832
- PMCID: PMC11449008
- DOI: 10.1093/neuonc/noae109
Super-enhancer-driven IRF2BP2 enhances ALK activity and promotes neuroblastoma cell proliferation
Abstract
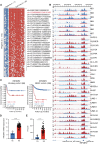
Background: Super-enhancers (SEs) typically govern the expression of critical oncogenes and play a fundamental role in the initiation and progression of cancer. Focusing on genes that are abnormally regulated by SE in cancer may be a new strategy for understanding pathogenesis. In the context of this investigation, we have identified a previously unreported SE-driven gene IRF2BP2 in neuroblastoma (NB).
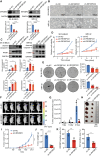
Methods: The expression and prognostic value of IRF2BP2 were detected in public databases and clinical samples. The effect of IRF2BP2 on NB cell growth and apoptosis was evaluated through in vivo and in vitro functional loss experiments. The molecular mechanism of IRF2BP2 was investigated by the study of chromatin regulatory regions and transcriptome sequencing.
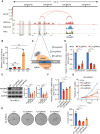
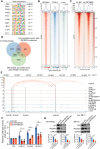
Results: The sustained high expression of IRF2BP2 results from the activation of a novel SE established by NB master transcription factors MYCN, MEIS2, and HAND2, and they form a new complex that regulates the gene network associated with the proliferation of NB cell populations. We also observed a significant enrichment of the AP-1 family at the binding sites of IRF2BP2. Remarkably, within NB cells, AP-1 plays a pivotal role in shaping the chromatin accessibility landscape, thereby exposing the binding site for IRF2BP2. This orchestrated action enables AP-1 and IRF2BP2 to collaboratively stimulate the expression of the NB susceptibility gene ALK, thereby upholding the highly proliferative phenotype characteristic of NB.
Conclusions: Our findings indicate that SE-driven IRF2BP2 can bind to AP-1 to maintain the survival of tumor cells via regulating chromatin accessibility of the NB susceptibility gene ALK.
Keywords: AP-1; IRF2BP2; chromatin accessibility; neuroblastoma; super-enhancer.
© The Author(s) 2024. Published by Oxford University Press on behalf of the Society for Neuro-Oncology. All rights reserved. For commercial re-use, please contact reprints@oup.com for reprints and translation rights for reprints. All other permissions can be obtained through our RightsLink service via the Permissions link on the article page on our site—for further information please contact journals.permissions@oup.com.
Conflict of interest statement
The authors have declared that no conflict of interest exists.
Figures



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

