SCC3 is an axial element essential for homologous chromosome pairing and synapsis
- PMID: 38864853
- PMCID: PMC11168746
- DOI: 10.7554/eLife.94180
SCC3 is an axial element essential for homologous chromosome pairing and synapsis
Abstract
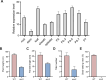
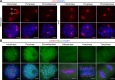
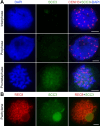
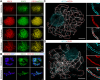
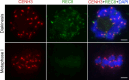
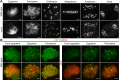
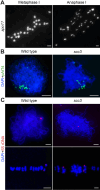
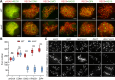
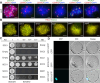
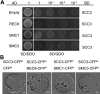
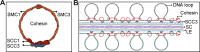
Cohesin is a multi-subunit protein that plays a pivotal role in holding sister chromatids together during cell division. Sister chromatid cohesion 3 (SCC3), constituents of cohesin complex, is highly conserved from yeast to mammals. Since the deletion of individual cohesin subunit always causes lethality, it is difficult to dissect its biological function in both mitosis and meiosis. Here, we obtained scc3 weak mutants using CRISPR-Cas9 system to explore its function during rice mitosis and meiosis. The scc3 weak mutants displayed obvious vegetative defects and complete sterility, underscoring the essential roles of SCC3 in both mitosis and meiosis. SCC3 is localized on chromatin from interphase to prometaphase in mitosis. However, in meiosis, SCC3 acts as an axial element during early prophase I and subsequently situates onto centromeric regions following the disassembly of the synaptonemal complex. The loading of SCC3 onto meiotic chromosomes depends on REC8. scc3 shows severe defects in homologous pairing and synapsis. Consequently, SCC3 functions as an axial element that is essential for maintaining homologous chromosome pairing and synapsis during meiosis.
Keywords: cell biology; genetics; genomics; meiocyte; meiosis; rice.
© 2024, Zhao, Ren, Zhao et al.
Conflict of interest statement
YZ, LR, TZ, HY, YM, HL, LC, BW, YS, YL, DT, ZC No competing interests declared
Figures







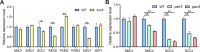
Update of
- doi: 10.1101/2023.11.22.568373
- doi: 10.7554/eLife.94180.1
- doi: 10.7554/eLife.94180.2
Similar articles
-
Meiosis-specific cohesin component, Stag3 is essential for maintaining centromere chromatid cohesion, and required for DNA repair and synapsis between homologous chromosomes.PLoS Genet. 2014 Jul 3;10(7):e1004413. doi: 10.1371/journal.pgen.1004413. eCollection 2014 Jul. PLoS Genet. 2014. PMID: 24992337 Free PMC article.
-
The cohesion protein SOLO associates with SMC1 and is required for synapsis, recombination, homolog bias and cohesion and pairing of centromeres in Drosophila Meiosis.PLoS Genet. 2013;9(7):e1003637. doi: 10.1371/journal.pgen.1003637. Epub 2013 Jul 18. PLoS Genet. 2013. PMID: 23874232 Free PMC article.
-
Genetic Interactions Between the Meiosis-Specific Cohesin Components, STAG3, REC8, and RAD21L.G3 (Bethesda). 2016 Jun 1;6(6):1713-24. doi: 10.1534/g3.116.029462. G3 (Bethesda). 2016. PMID: 27172213 Free PMC article.
-
Rec8 Cohesin: A Structural Platform for Shaping the Meiotic Chromosomes.Genes (Basel). 2022 Jan 22;13(2):200. doi: 10.3390/genes13020200. Genes (Basel). 2022. PMID: 35205245 Free PMC article. Review.
-
The cohesin complex in mammalian meiosis.Genes Cells. 2019 Jan;24(1):6-30. doi: 10.1111/gtc.12652. Epub 2018 Nov 27. Genes Cells. 2019. PMID: 30479058 Free PMC article. Review.
References
-
- Agostinho A, Manneberg O, van Schendel R, Hernández-Hernández A, Kouznetsova A, Blom H, Brismar H, Höög C. High density of REC8 constrains sister chromatid axes and prevents illegitimate synaptonemal complex formation. EMBO Reports. 2016;17:901–913. doi: 10.15252/embr.201642030. - DOI - PMC - PubMed
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources

