AI analysis of super-resolution microscopy: Biological discovery in the absence of ground truth
- PMID: 38865088
- PMCID: PMC11169916
- DOI: 10.1083/jcb.202311073
AI analysis of super-resolution microscopy: Biological discovery in the absence of ground truth
Abstract
Super-resolution microscopy, or nanoscopy, enables the use of fluorescent-based molecular localization tools to study molecular structure at the nanoscale level in the intact cell, bridging the mesoscale gap to classical structural biology methodologies. Analysis of super-resolution data by artificial intelligence (AI), such as machine learning, offers tremendous potential for the discovery of new biology, that, by definition, is not known and lacks ground truth. Herein, we describe the application of weakly supervised paradigms to super-resolution microscopy and its potential to enable the accelerated exploration of the nanoscale architecture of subcellular macromolecules and organelles.
© 2024 Nabi et al.
Conflict of interest statement
Disclosures: All authors have completed and submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest. I. Nabi reported a patent to WO/2019/109181 issued “UBC/SFU.” G. Hamarneh reported a patent to WO/2019/109181 issued “UBC & SFU.” No other disclosures were reported.
Figures



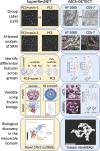
Similar articles
-
Self-inspired learning for denoising live-cell super-resolution microscopy.Nat Methods. 2024 Oct;21(10):1895-1908. doi: 10.1038/s41592-024-02400-9. Epub 2024 Sep 11. Nat Methods. 2024. PMID: 39261639
-
Nanoscale single-vesicle analysis: High-throughput approaches through AI-enhanced super-resolution image analysis.Biosens Bioelectron. 2024 Nov 1;263:116629. doi: 10.1016/j.bios.2024.116629. Epub 2024 Aug 5. Biosens Bioelectron. 2024. PMID: 39106689
-
Enhanced super-resolution microscopy by extreme value based emitter recovery.Sci Rep. 2021 Oct 14;11(1):20417. doi: 10.1038/s41598-021-00066-3. Sci Rep. 2021. PMID: 34650088 Free PMC article.
-
Machine learning-enabled computer vision for plant phenotyping: a primer on AI/ML and a case study on stomatal patterning.J Exp Bot. 2024 Nov 15;75(21):6683-6703. doi: 10.1093/jxb/erae395. J Exp Bot. 2024. PMID: 39363775 Free PMC article. Review.
-
Three-Dimensional Single-Molecule Localization Microscopy in Whole-Cell and Tissue Specimens.Annu Rev Biomed Eng. 2020 Jun 4;22:155-184. doi: 10.1146/annurev-bioeng-060418-052203. Epub 2020 Apr 3. Annu Rev Biomed Eng. 2020. PMID: 32243765 Free PMC article. Review.
Cited by
-
An update on recent advances in fluorescent materials for fluorescence molecular imaging: a review.RSC Adv. 2025 Jun 30;15(28):22267-22284. doi: 10.1039/d5ra03102h. eCollection 2025 Jun 30. RSC Adv. 2025. PMID: 40599579 Free PMC article. Review.
-
Comparative Analysis of SPLICS and MCS-DETECT for Detecting Mitochondria-ER Contact Sites (MERCs).Contact (Thousand Oaks). 2025 Mar 18;8:25152564251313721. doi: 10.1177/25152564251313721. eCollection 2025 Jan-Dec. Contact (Thousand Oaks). 2025. PMID: 40115170 Free PMC article.
-
Should Artificial Intelligence Play a Durable Role in Biomedical Research and Practice?Int J Mol Sci. 2024 Dec 13;25(24):13371. doi: 10.3390/ijms252413371. Int J Mol Sci. 2024. PMID: 39769135 Free PMC article. Review.
-
Closing the multichannel gap through computational reconstruction of interaction in super-resolution microscopy.Patterns (N Y). 2025 Mar 27;6(5):101181. doi: 10.1016/j.patter.2025.101181. eCollection 2025 May 9. Patterns (N Y). 2025. PMID: 40486966 Free PMC article. Review.
-
Scaffolds and the scaffolding domain: an alternative paradigm for caveolin-1 signaling.Biochem Soc Trans. 2024 Apr 24;52(2):947-959. doi: 10.1042/BST20231570. Biochem Soc Trans. 2024. PMID: 38526159 Free PMC article. Review.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

