De novo TLK1 and MDM1 mutations in a patient with a neurodevelopmental disorder and immunodeficiency
- PMID: 38868186
- PMCID: PMC11166698
- DOI: 10.1016/j.isci.2024.109984
De novo TLK1 and MDM1 mutations in a patient with a neurodevelopmental disorder and immunodeficiency
Abstract
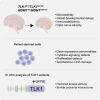
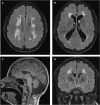
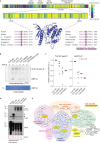
The Tousled-like kinases 1 and 2 (TLK1/TLK2) regulate DNA replication, repair and chromatin maintenance. TLK2 variants underlie the neurodevelopmental disorder (NDD) 'Intellectual Disability, Autosomal Dominant 57' (MRD57), characterized by intellectual disability and microcephaly. Several TLK1 variants have been reported in NDDs but their functional significance is unknown. A male patient presenting with ID, seizures, global developmental delay, hypothyroidism, and primary immunodeficiency was determined to have a heterozygous TLK1 variant (c.1435C>G, p.Q479E), as well as a mutation in MDM1 (c.1197dupT, p.K400∗). Cells expressing TLK1 p.Q479E exhibited reduced cytokine responses and elevated DNA damage, but not increased radiation sensitivity or DNA repair defects. The TLK1 p.Q479E variant impaired kinase activity but not proximal protein interactions. Our study provides the first functional characterization of NDD-associated TLK1 variants and suggests that, such as TLK2, TLK1 variants may impact development in multiple tissues and should be considered in the diagnosis of rare NDDs.
Keywords: genetics; immunology; neuroscience.
Published by Elsevier Inc.
Conflict of interest statement
The authors declare no competing interests.
Figures



Update of
-
Identification of a de novo mutation in TLK1 associated with a neurodevelopmental disorder and immunodeficiency.medRxiv [Preprint]. 2023 Aug 24:2023.08.22.23294267. doi: 10.1101/2023.08.22.23294267. medRxiv. 2023. Update in: iScience. 2024 May 18;27(6):109984. doi: 10.1016/j.isci.2024.109984. PMID: 37662408 Free PMC article. Updated. Preprint.
References
-
- Moshkin Y.M., Kan T.W., Goodfellow H., Bezstarosti K., Maeda R.K., Pilyugin M., Karch F., Bray S.J., Demmers J.A.A., Verrijzer C.P. Histone chaperones ASF1 and NAP1 differentially modulate removal of active histone marks by LID-RPD3 complexes during NOTCH silencing. Mol. Cell. 2009;35:782–793. doi: 10.1016/j.molcel.2009.07.020. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

