Gut microbiome encoded purine and amino acid pathways present prospective biomarkers for predicting metformin therapy efficacy in newly diagnosed T2D patients
- PMID: 38868903
- PMCID: PMC11178274
- DOI: 10.1080/19490976.2024.2361491
Gut microbiome encoded purine and amino acid pathways present prospective biomarkers for predicting metformin therapy efficacy in newly diagnosed T2D patients
Abstract
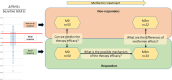
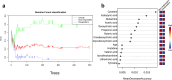
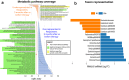
Metformin is widely used for treating type 2 diabetes mellitus (T2D). However, the efficacy of metformin monotherapy is highly variable within the human population. Understanding the potential indirect or synergistic effects of metformin on gut microbiota composition and encoded functions could potentially offer new insights into predicting treatment efficacy and designing more personalized treatments in the future. We combined targeted metabolomics and metagenomic profiling of gut microbiomes in newly diagnosed T2D patients before and after metformin therapy to identify potential pre-treatment biomarkers and functional signatures for metformin efficacy and induced changes in metformin therapy responders. Our sequencing data were largely corroborated by our metabolic profiling and identified that pre-treatment enrichment of gut microbial functions encoding purine degradation and glutamate biosynthesis was associated with good therapy response. Furthermore, we identified changes in glutamine-associated amino acid (arginine, ornithine, putrescine) metabolism that characterize differences in metformin efficacy before and after the therapy. Moreover, metformin Responders' microbiota displayed a shifted balance between bacterial lipidA synthesis and degradation as well as alterations in glutamate-dependent metabolism of N-acetyl-galactosamine and its derivatives (e.g. CMP-pseudaminate) which suggest potential modulation of bacterial cell walls and human gut barrier, thus mediating changes in microbiome composition. Together, our data suggest that glutamine and associated amino acid metabolism as well as purine degradation products may potentially condition metformin activity via its multiple effects on microbiome functional composition and therefore serve as important biomarkers for predicting metformin efficacy.
Keywords: Gut microbiome; T2D; biomarkers; functional profile; metabolic analysis; metformin.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures







Similar articles
-
Metformin-induced changes in the gut microbiome and plasma metabolome are associated with cognition in men.Metabolism. 2024 Aug;157:155941. doi: 10.1016/j.metabol.2024.155941. Epub 2024 Jun 12. Metabolism. 2024. PMID: 38871078 Clinical Trial.
-
A specific gut microbiota and metabolomic profiles shifts related to antidiabetic action: The similar and complementary antidiabetic properties of type 3 resistant starch from Canna edulis and metformin.Pharmacol Res. 2020 Sep;159:104985. doi: 10.1016/j.phrs.2020.104985. Epub 2020 Jun 3. Pharmacol Res. 2020. PMID: 32504839
-
Interplay between gut microbiota and tryptophan metabolism in type 2 diabetic mice treated with metformin.Microbiol Spectr. 2024 Oct 3;12(10):e0029124. doi: 10.1128/spectrum.00291-24. Epub 2024 Aug 20. Microbiol Spectr. 2024. PMID: 39162538 Free PMC article.
-
Metformin: old friend, new ways of action-implication of the gut microbiome?Curr Opin Clin Nutr Metab Care. 2018 Jul;21(4):294-301. doi: 10.1097/MCO.0000000000000468. Curr Opin Clin Nutr Metab Care. 2018. PMID: 29634493 Review.
-
The Relationship between the Gut Microbiome and Metformin as a Key for Treating Type 2 Diabetes Mellitus.Int J Mol Sci. 2021 Mar 30;22(7):3566. doi: 10.3390/ijms22073566. Int J Mol Sci. 2021. PMID: 33808194 Free PMC article. Review.
Cited by
-
Aqueous humor metabolomic profiling identifies a distinct signature in pseudoexfoliation syndrome.Front Mol Biosci. 2025 Jan 23;11:1487115. doi: 10.3389/fmolb.2024.1487115. eCollection 2024. Front Mol Biosci. 2025. PMID: 39917180 Free PMC article.
-
Lactobacillus murinus alleviates insulin resistance via promoting L-citrulline synthesis.J Endocrinol Invest. 2025 Apr;48(4):1005-1015. doi: 10.1007/s40618-024-02500-5. Epub 2024 Nov 19. J Endocrinol Invest. 2025. PMID: 39560906
-
Specific microbial ratio in the gut microbiome is associated with multiple sclerosis.Proc Natl Acad Sci U S A. 2025 Mar 11;122(10):e2413953122. doi: 10.1073/pnas.2413953122. Epub 2025 Mar 3. Proc Natl Acad Sci U S A. 2025. PMID: 40030030
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
