Deep Amplicon Sequencing Reveals Culture-dependent Clonal Selection of Mycobacterium tuberculosis in Clinical Samples
- PMID: 38870522
- PMCID: PMC11978391
- DOI: 10.1093/gpbjnl/qzae046
Deep Amplicon Sequencing Reveals Culture-dependent Clonal Selection of Mycobacterium tuberculosis in Clinical Samples
Abstract
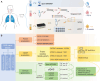
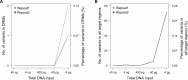
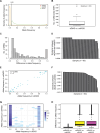
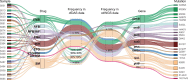
The commonly-used drug susceptibility testing (DST) relies on bacterial culture and faces shortcomings such as long turnaround time and clonal/subclonal selection biases. Here, we developed a targeted deep amplicon sequencing (DAS) method directly applied to clinical specimens. In this DAS panel, we examined 941 drug-resistant mutations (DRMs) associated with 20 anti-tuberculosis drugs with only 4 pg of initial DNA input, and reduced the clinical testing time from 20 days to 2 days. A prospective study was conducted using 115 clinical specimens, predominantly positive for the Xpert® Mycobacterium tuberculosis/rifampicin (Xpert MTB/RIF) assay, to evaluate DRM detection. DAS was performed on culture-free specimens, while culture-dependent isolates were used for phenotypic DST, DAS, and whole-genome sequencing (WGS). For in silico molecular DST, our result based on DAS panel revealed the similar accuracy to three published reports based on WGS. For 82 isolates, application of DAS using the resistance-determining mutation method showed better accuracy (93.03% vs. 92.16%), sensitivity (96.10% vs. 95.02%), and specificity (91.33% vs. 90.62%) than WGS using the Mykrobe software. Compared to culture-dependent WGS, culture-free DAS provides a full picture of sequence variation at the population level, exhibiting in detail the gain-and-loss variants caused by bacterial culture. Our study performs a systematic verification of the advantages of DAS in clinical applications and comprehensively illustrates the discrepancies in Mycobacterium tuberculosis before and after culture.
Keywords: Mycobacterium tuberculosis; Culture-dependent; Culture-free; Deep amplicon sequencing; Drug susceptibility testing.
© The Author(s) 2024. Published by Oxford University Press and Science Press on behalf of the Beijing Institute of Genomics, Chinese Academy of Sciences / China National Center for Bioinformation and Genetics Society of China.
Conflict of interest statement
The authors have declared no competing interests.
Figures
Similar articles
-
Unveiling the complexity of rifampicin drug susceptibility testing in Mycobacterium tuberculosis: comparative analysis with next-generation sequencing.J Med Microbiol. 2024 Sep;73(9). doi: 10.1099/jmm.0.001884. J Med Microbiol. 2024. PMID: 39229883
-
Accuracy of whole genome sequencing versus phenotypic (MGIT) and commercial molecular tests for detection of drug-resistant Mycobacterium tuberculosis isolated from patients in Brazil and Mozambique.Tuberculosis (Edinb). 2018 May;110:59-67. doi: 10.1016/j.tube.2018.04.003. Epub 2018 Apr 5. Tuberculosis (Edinb). 2018. PMID: 29779775 Free PMC article.
-
Comparison of targeted next-generation sequencing and the Xpert MTB/RIF assay for detection of Mycobacterium tuberculosis in clinical isolates and sputum specimens.Microbiol Spectr. 2024 May 2;12(5):e0409823. doi: 10.1128/spectrum.04098-23. Epub 2024 Apr 11. Microbiol Spectr. 2024. PMID: 38602399 Free PMC article.
-
Xpert MTB/RIF Ultra and Xpert MTB/RIF assays for extrapulmonary tuberculosis and rifampicin resistance in adults.Cochrane Database Syst Rev. 2021 Jan 15;1(1):CD012768. doi: 10.1002/14651858.CD012768.pub3. Cochrane Database Syst Rev. 2021. PMID: 33448348 Free PMC article.
-
Xpert MTB/RIF and Xpert MTB/RIF Ultra assays for active tuberculosis and rifampicin resistance in children.Cochrane Database Syst Rev. 2020 Aug 27;8(8):CD013359. doi: 10.1002/14651858.CD013359.pub2. Cochrane Database Syst Rev. 2020. Update in: Cochrane Database Syst Rev. 2022 Sep 6;9:CD013359. doi: 10.1002/14651858.CD013359.pub3. PMID: 32853411 Free PMC article. Updated.
Cited by
-
Effect of Bdq-Containing Regimen and Molecular Detection of Bdq Resistance among Pre-XDR-TB Patients with Unfavorable Outcomes.Int J Gen Med. 2025 Aug 14;18:4469-4480. doi: 10.2147/IJGM.S538660. eCollection 2025. Int J Gen Med. 2025. PMID: 40837941 Free PMC article.
-
Whole-genome sequencing reveals transmission pattern and drug resistance of Mycobacterium tuberculosis intra- or inter-hosts.Front Cell Infect Microbiol. 2025 Jan 21;14:1488547. doi: 10.3389/fcimb.2024.1488547. eCollection 2024. Front Cell Infect Microbiol. 2025. PMID: 39906217 Free PMC article.
References
-
- World Health Organization. Global tuberculosis report 2021. [Internet]. Geneva, Switzerland: World Health Organization; 2021, https://www.who.int/publications/digital/global-tuberculosis-report-2021.
-
- Nikolayevskyy V, Kranzer K, Niemann S, Drobniewski F. Whole genome sequencing of Mycobacterium tuberculosis for detection of recent transmission and tracing outbreaks: a systematic review. Tuberculosis 2016;98:77–85. - PubMed
-
- Goig GA, Cancino-Munoz I, Torres-Puente M, Villamayor LM, Navarro D, Borras R, et al. Whole-genome sequencing of Mycobacterium tuberculosis directly from clinical samples for high-resolution genomic epidemiology and drug resistance surveillance: an observational study. Lancet Microbe 2020;1:e175–83. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

