Supervised learning of enhancer-promoter specificity based on genome-wide perturbation studies highlights areas for improvement in learning
- PMID: 38870532
- PMCID: PMC11211214
- DOI: 10.1093/bioinformatics/btae367
Supervised learning of enhancer-promoter specificity based on genome-wide perturbation studies highlights areas for improvement in learning
Abstract
Motivation: Understanding the rules that govern enhancer-driven transcription remains a central unsolved problem in genomics. Now with multiple massively parallel enhancer perturbation assays published, there are enough data that we can utilize to learn to predict enhancer-promoter (EP) relationships in a data-driven manner.
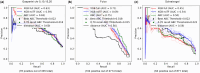
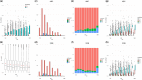
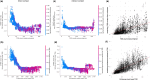
Results: We applied machine learning to one of the largest enhancer perturbation studies integrated with transcription factor (TF) and histone modification ChIP-seq. The results uncovered a discrepancy in the prediction of genome-wide data compared to data from targeted experiments. Relative strength of contact was important for prediction, confirming the basic principle of EP regulation. Novel features such as the density of the enhancers/promoters in the genomic region was found to be important, highlighting our lack of understanding on how other elements in the region contribute to the regulation. Several TF peaks were identified that improved the prediction by identifying the negatives and reducing False Positives. In summary, integrating genomic assays with enhancer perturbation studies increased the accuracy of the model, and provided novel insights into the understanding of enhancer-driven transcription.
Availability and implementation: The trained models, data, and the source code are available at http://doi.org/10.5281/zenodo.11290386 and https://github.com/HanLabUNLV/sleps.
© The Author(s) 2024. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures




Similar articles
-
Enhancer prediction in the human genome by probabilistic modelling of the chromatin feature patterns.BMC Bioinformatics. 2020 Jul 20;21(1):317. doi: 10.1186/s12859-020-03621-3. BMC Bioinformatics. 2020. PMID: 32689977 Free PMC article.
-
Enhancer identification in mouse embryonic stem cells using integrative modeling of chromatin and genomic features.BMC Genomics. 2012 Apr 26;13:152. doi: 10.1186/1471-2164-13-152. BMC Genomics. 2012. PMID: 22537144 Free PMC article.
-
Predicting enhancers in mammalian genomes using supervised hidden Markov models.BMC Bioinformatics. 2019 Mar 27;20(1):157. doi: 10.1186/s12859-019-2708-6. BMC Bioinformatics. 2019. PMID: 30917778 Free PMC article.
-
Enhancer target prediction: state-of-the-art approaches and future prospects.Biochem Soc Trans. 2023 Oct 31;51(5):1975-1988. doi: 10.1042/BST20230917. Biochem Soc Trans. 2023. PMID: 37830459 Review.
-
Decoding enhancers using massively parallel reporter assays.Genomics. 2015 Sep;106(3):159-164. doi: 10.1016/j.ygeno.2015.06.005. Epub 2015 Jun 10. Genomics. 2015. PMID: 26072433 Free PMC article. Review.
References
-
- Akiba T, Sano S, Yanase T. et al. Optuna: a next-generation hyperparameter optimization framework. In, Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, KDD ’19. Anchorage AK USA. Association for Computing Machinery, New York, NY, USA, pp. 2623–2631.
-
- Cao Q, Anyansi C, Hu X. et al. Reconstruction of enhancer–target networks in 935 samples of human primary cells, tissues and cell lines. Nat Genet 2017;49:1428–36. - PubMed
-
- Chen T, Guestrin C. XGBoost: a scalable tree boosting system. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco California USA. 2016, New York, NY, USA: Association for Computing Machinery, pp. 2623–2631.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

