Ischemic Stroke with Comorbid Cancer Has Specific miRNA-mRNA Networks in Blood That Vary by Ischemic Stroke Mechanism
- PMID: 38874304
- PMCID: PMC11849972
- DOI: 10.1002/ana.26997
Ischemic Stroke with Comorbid Cancer Has Specific miRNA-mRNA Networks in Blood That Vary by Ischemic Stroke Mechanism
Abstract
Objective: Approximately half of ischemic strokes (IS) in cancer patients are cryptogenic, with many presumed cardioembolic. We evaluated whether there were specific miRNA and mRNA transcriptome architectures in peripheral blood of IS patients with and without comorbid cancer, and between cardioembolic versus noncardioembolic IS etiologies in comorbid cancer.
Methods: We studied patients with cancer and IS (CS; n = 42), stroke only (SO; n = 41), and cancer only (n = 28), and vascular risk factor-matched controls (n = 30). mRNA-Seq and miRNA-Seq data, analyzed with linear regression models, identified differentially expressed genes in CS versus SO and in cardioembolic versus noncardioembolic CS, and miRNA-mRNA regulatory pairs. Network-level analyses identified stroke etiology-specific responses in CS.
Results: A total of 2,085 mRNAs and 31 miRNAs were differentially expressed between CS and SO. In CS, 122 and 35 miRNA-mRNA regulatory pairs, and 5 and 3 coexpressed gene modules, were associated with cardioembolic and noncardioembolic CS, respectively. Complement, growth factor, and immune/inflammatory pathways showed differences between IS etiologies in CS. A 15-gene biomarker panel assembled from a derivation cohort (n = 50) correctly classified 81% of CS and 71% of SO participants in a validation cohort (n = 33). Another 15-gene panel correctly identified etiologies for 13 of 13 CS-cardioembolic and 11 of 11 CS-noncardioembolic participants upon cross-validation; 11 of 16 CS-cryptogenic participants were predicted cardioembolic.
Interpretation: We discovered unique mRNA and miRNA transcriptome architecture in CS and SO, and in CS with different IS etiologies. Cardioembolic and noncardioembolic etiologies in CS showed unique coexpression networks and potential master regulators. These may help distinguish CS from SO and identify IS etiology in cryptogenic CS patients. ANN NEUROL 2024;96:565-581.
© 2024 The Author(s). Annals of Neurology published by Wiley Periodicals LLC on behalf of American Neurological Association.
Conflict of interest statement
Potential Conflicts of Interest
G.J., F.R.S., and B.S. hold a patent on gene expression signatures for predicting stroke and its causes. All other authors report no competing interests.
Figures



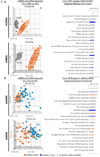






References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

