Cellular endosomal potassium ion flux regulates arenavirus uncoating during virus entry
- PMID: 38874413
- PMCID: PMC11253613
- DOI: 10.1128/mbio.01684-23
Cellular endosomal potassium ion flux regulates arenavirus uncoating during virus entry
Abstract
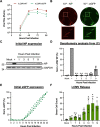
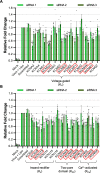
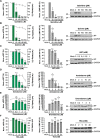
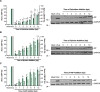
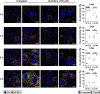
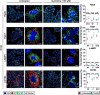
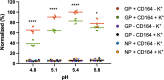
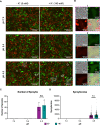
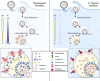
Lymphocytic choriomeningitis virus (LCMV) is an enveloped and segmented negative-sense RNA virus classified within the Arenaviridae family of the Bunyavirales order. LCMV is associated with fatal disease in immunocompromised populations and, as the prototypical arenavirus member, acts as a model for the many highly pathogenic members of the Arenaviridae family, such as Junín, Lassa, and Lujo viruses, all of which are associated with devastating hemorrhagic fevers. To enter cells, the LCMV envelope fuses with late endosomal membranes, for which two established requirements are low pH and interaction between the LCMV glycoprotein (GP) spike and secondary receptor CD164. LCMV subsequently uncoats, where the RNA genome-associated nucleoprotein (NP) separates from the Z protein matrix layer, releasing the viral genome into the cytosol. To further examine LCMV endosome escape, we performed an siRNA screen which identified host cell potassium ion (K+) channels as important for LCMV infection, with pharmacological inhibition confirming K+ channel involvement during the LCMV entry phase completely abrogating productive infection. To better understand the K+-mediated block in infection, we tracked incoming virions along their entry pathway under physiological conditions, where uncoating was signified by separation of NP and Z proteins. In contrast, K+ channel blockade prevented uncoating, trapping virions within Rab7 and CD164-positive endosomes, identifying K+ as a third LCMV entry requirement. K+ did not increase GP-CD164 binding or alter GP-CD164-dependent fusion. Thus, we propose that K+ mediates uncoating by modulating NP-Z interactions within the virion interior. These results suggest K+ channels represent a potential anti-arenaviral target.IMPORTANCEArenaviruses can cause fatal human disease for which approved preventative or therapeutic options are not available. Here, using the prototypical LCMV, we identified K+ channels as critical for arenavirus infection, playing a vital role during the entry phase of the infection cycle. We showed that blocking K+ channel function resulted in entrapment of LCMV particles within late endosomal compartments, thus preventing productive replication. Our data suggest K+ is required for LCMV uncoating and genome release by modulating interactions between the viral nucleoprotein and the matrix protein layer inside the virus particle.
Keywords: LCMV; arenavirus; ion channels; potassium.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Lymphocytic choriomeningitis arenavirus requires cellular COPI and AP-4 complexes for efficient virion production.J Virol. 2024 Mar 19;98(3):e0200623. doi: 10.1128/jvi.02006-23. Epub 2024 Feb 9. J Virol. 2024. PMID: 38334330 Free PMC article.
-
Old world arenaviruses enter the host cell via the multivesicular body and depend on the endosomal sorting complex required for transport.PLoS Pathog. 2011 Sep;7(9):e1002232. doi: 10.1371/journal.ppat.1002232. Epub 2011 Sep 8. PLoS Pathog. 2011. PMID: 21931550 Free PMC article.
-
Different mechanisms of cell entry by human-pathogenic Old World and New World arenaviruses.J Virol. 2008 Aug;82(15):7677-87. doi: 10.1128/JVI.00560-08. Epub 2008 May 28. J Virol. 2008. PMID: 18508885 Free PMC article.
-
Receptor binding and cell entry of Old World arenaviruses reveal novel aspects of virus-host interaction.Virology. 2009 May 10;387(2):245-9. doi: 10.1016/j.virol.2009.02.042. Epub 2009 Mar 25. Virology. 2009. PMID: 19324387 Review.
-
Arenavirus Quasispecies and Their Biological Implications.Curr Top Microbiol Immunol. 2016;392:231-76. doi: 10.1007/82_2015_468. Curr Top Microbiol Immunol. 2016. PMID: 26472215 Free PMC article. Review.
Cited by
-
Lymphocytic choriomeningitis arenavirus requires cellular COPI and AP-4 complexes for efficient virion production.J Virol. 2024 Mar 19;98(3):e0200623. doi: 10.1128/jvi.02006-23. Epub 2024 Feb 9. J Virol. 2024. PMID: 38334330 Free PMC article.
References
-
- Walker PJ, Siddell SG, Lefkowitz EJ, Mushegian AR, Dempsey DM, Dutilh BE, Harrach B, Harrison RL, Hendrickson RC, Junglen S, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Nibert M, Rubino L, Sabanadzovic S, Simmonds P, Varsani A, Zerbini FM, Davison AJ. 2019. Changes to virus taxonomy and the International code of virus classification and nomenclature ratified by the International committee on taxonomy of viruses. Arch Virol 164:2417–2429. doi:10.1007/s00705-019-04306-w - DOI - PubMed
-
- Radoshitzky SR, Buchmeier MJ, Charrel RN, Clegg JCS, Gonzalez J-P, Günther S, Hepojoki J, Kuhn JH, Lukashevich IS, Romanowski V, Salvato MS, Sironi M, Stenglein MD, de la Torre JC, Ictv Report Consortium . 2019. ICTV virus taxonomy profile: arenaviridae. J Gen Virol 100:1200–1201. doi:10.1099/jgv.0.001280 - DOI - PubMed
-
- Kernéis S, Koivogui L, Magassouba N, Koulemou K, Lewis R, Aplogan A, Grais RF, Guerin PJ, Fichet-Calvet E. 2009. Prevalence and risk factors of Lassa seropositivity in inhabitants of the forest region of guinea: a cross-sectional study. PLoS Negl Trop Dis 3:e548. doi:10.1371/journal.pntd.0000548 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

