Cadherin-11 contributes to the heterogenous and dynamic Wnt-Wnt-β-catenin pathway activation in Ewing sarcoma
- PMID: 38875295
- PMCID: PMC11178195
- DOI: 10.1371/journal.pone.0305490
Cadherin-11 contributes to the heterogenous and dynamic Wnt-Wnt-β-catenin pathway activation in Ewing sarcoma
Abstract
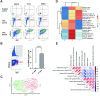
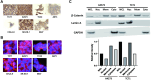
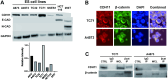
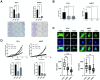
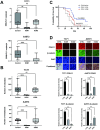
Ewing sarcoma is the second most common bone cancer in children, and while patients who present with metastatic disease at the time of diagnosis have a dismal prognosis. Ewing sarcoma tumors are driven by the fusion gene EWS/Fli1, and while these tumors are genetically homogenous, the transcriptional heterogeneity can lead to a variety of cellular processes including metastasis. In this study, we demonstrate that in Ewing sarcoma cells, the canonical Wnt/β-Catenin signaling pathway is heterogeneously activated in vitro and in vivo, correlating with hypoxia and EWS/Fli1 activity. Ewing sarcoma cells predominantly express β-Catenin on the cell membrane bound to CDH11, which can respond to exogenous Wnt ligands leading to the immediate activation of Wnt/β-Catenin signaling within a tumor. Knockdown of CDH11 leads to delayed and decreased response to exogenous Wnt ligand stimulation, and ultimately decreased metastatic propensity. Our findings strongly indicate that CDH11 is a key component of regulating Wnt//β-Catenin signaling heterogeneity within Ewing sarcoma tumors, and is a promising molecular target to alter Wnt//β-Catenin signaling in Ewing sarcoma patients.
Copyright: © 2024 Shirai et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Activation of Wnt/β-Catenin in Ewing Sarcoma Cells Antagonizes EWS/ETS Function and Promotes Phenotypic Transition to More Metastatic Cell States.Cancer Res. 2016 Sep 1;76(17):5040-53. doi: 10.1158/0008-5472.CAN-15-3422. Epub 2016 Jun 30. Cancer Res. 2016. PMID: 27364557 Free PMC article.
-
Wnt/β-catenin-activated Ewing sarcoma cells promote the angiogenic switch.JCI Insight. 2020 Jul 9;5(13):e135188. doi: 10.1172/jci.insight.135188. JCI Insight. 2020. PMID: 32544094 Free PMC article.
-
EWS-FLI1 and Menin Converge to Regulate ATF4 Activity in Ewing Sarcoma.Mol Cancer Res. 2021 Jul;19(7):1182-1195. doi: 10.1158/1541-7786.MCR-20-0679. Epub 2021 Mar 19. Mol Cancer Res. 2021. PMID: 33741715 Free PMC article.
-
New Insights about the Wnt/β-Catenin Signaling Pathway in Primary Bone Tumors and Their Microenvironment: A Promising Target to Develop Therapeutic Strategies?Int J Mol Sci. 2019 Jul 31;20(15):3751. doi: 10.3390/ijms20153751. Int J Mol Sci. 2019. PMID: 31370265 Free PMC article. Review.
-
Molecular pathogenesis of Ewing sarcoma: new therapeutic and transcriptional targets.Annu Rev Pathol. 2012;7:145-59. doi: 10.1146/annurev-pathol-011110-130237. Epub 2011 Sep 19. Annu Rev Pathol. 2012. PMID: 21942527 Free PMC article. Review.
References
-
- Womer RB, West DC, Krailo MD, Dickman PS, Pawel BR, Grier HE, et al.. Randomized controlled trial of interval-compressed chemotherapy for the treatment of localized Ewing sarcoma: a report from the Children’s Oncology Group. J Clin Oncol. 2012;30(33):4148–54. Epub 20121022. doi: 10.1200/JCO.2011.41.5703 ; PubMed Central PMCID: PMC3494838. - DOI - PMC - PubMed
-
- Rosen G, Wollner N, Tan C, Wu SJ, Hajdu SI, Cham W, et al.. Proceedings: Disease-free survival in children with Ewing’s sarcoma treated with radiation therapy and adjuvant four-drug sequential chemotherapy. Cancer. 1974;33(2):384–93. Epub 1974/02/01. doi: 10.1002/1097-0142(197402)33:2<384::aid-cncr2820330213>3.0.co;2-t . - DOI - PubMed
-
- Nesbit ME Jr, Gehan EA, Burgert EO Jr., Vietti TJ, Cangir A, Tefft M, et al.. Multimodal therapy for the management of primary, nonmetastatic Ewing’s sarcoma of bone: a long-term follow-up of the First Intergroup study. J Clin Oncol. 1990;8(10):1664–74. Epub 1990/10/01. doi: 10.1200/JCO.1990.8.10.1664 . - DOI - PubMed
-
- Tirode F, Surdez D, Ma X, Parker M, Le Deley MC, Bahrami A, et al.. Genomic landscape of Ewing sarcoma defines an aggressive subtype with co-association of STAG2 and TP53 mutations. Cancer Discov. 2014;4(11):1342–53. Epub 20140915. doi: 10.1158/2159-8290.CD-14-0622 ; PubMed Central PMCID: PMC4264969. - DOI - PMC - PubMed
-
- Crompton BD, Stewart C, Taylor-Weiner A, Alexe G, Kurek KC, Calicchio ML, et al.. The genomic landscape of pediatric Ewing sarcoma. Cancer Discov. 2014;4(11):1326–41. Epub 2014/09/05. doi: 10.1158/2159-8290.CD-13-1037 . - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

