The pigeon circovirus evolution, epidemiology and interaction with the host immune system under One Loft Race rearing conditions
- PMID: 38877168
- PMCID: PMC11178769
- DOI: 10.1038/s41598-024-64587-3
The pigeon circovirus evolution, epidemiology and interaction with the host immune system under One Loft Race rearing conditions
Abstract
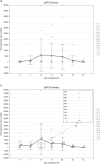
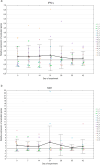
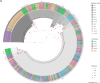
This study was aimed to investigate the frequency of PiCV recombination, the kinetics of PiCV viremia and shedding and the correlation between viral replication and host immune response in young pigeons subclinically infected with various PiCV variants and kept under conditions mimicking the OLR system. Fifteen racing pigeons originating from five breeding facilities were housed together for six weeks. Blood and cloacal swab samples were collected from birds every seven days to recover complete PiCV genomes and determine PiCV genetic diversity and recombination dynamics, as well as to assess virus shedding rate, level of viremia, expression of selected genes and level of anti-PiCV antibodies. Three hundred and eighty-eight complete PiCV genomes were obtained and thirteen genotypes were distinguished. Twenty-five recombination events were detected. Recombinants emerged during the first three weeks of the experiment which was consistent with the peak level of viremia and viral shedding. A further decrease in viremia and shedding partially corresponded with IFN-γ and MX1 gene expression and antibody dynamics. Considering the role of OLR pigeon rearing system in spreading infectious agents and allowing their recombination, it would be reasonable to reflect on the relevance of pigeon racing from both an animal welfare and epidemiological perspective.
Keywords: Gene expression; One Loft Race; Pigeon circovirus; Recombination; Viral evolution; ddPCR.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
A Pilot Study Investigating the Dynamics of Pigeon Circovirus Recombination in Domesticated Pigeons Housed in a Single Loft.Viruses. 2021 May 22;13(6):964. doi: 10.3390/v13060964. Viruses. 2021. PMID: 34067378 Free PMC article.
-
Pigeon circoviruses from feral pigeons in Australia demonstrate extensive recombination and genetic admixture with other circoviruses.Avian Pathol. 2019 Dec;48(6):512-520. doi: 10.1080/03079457.2019.1629391. Epub 2019 Jul 11. Avian Pathol. 2019. PMID: 31199167
-
Influence of pigeon interferon alpha (PiIFN-α) on pigeon circovirus (PiCV) replication and cytokine expression in Columba livia.Vet Microbiol. 2020 Mar;242:108591. doi: 10.1016/j.vetmic.2020.108591. Epub 2020 Jan 24. Vet Microbiol. 2020. PMID: 32122595
-
The epidemiology, molecular characterization and clinical pathology of circovirus infections in pigeons - current knowledge.Vet Q. 2017 Dec;37(1):166-174. doi: 10.1080/01652176.2017.1325972. Vet Q. 2017. PMID: 28463055 Review.
-
Pigeon Circovirus over Three Decades of Research: Bibliometrics, Scoping Review, and Perspectives.Viruses. 2022 Jul 8;14(7):1498. doi: 10.3390/v14071498. Viruses. 2022. PMID: 35891478 Free PMC article.
Cited by
-
Assessing reliability and accuracy of qPCR, dPCR and ddPCR for estimating mitochondrial DNA copy number in songbird blood and sperm cells.PeerJ. 2025 Apr 11;13:e19278. doi: 10.7717/peerj.19278. eCollection 2025. PeerJ. 2025. PMID: 40231068 Free PMC article.
-
Epidemiological and evolutionary analysis of canine circovirus from 1996 to 2023.BMC Vet Res. 2024 Jul 20;20(1):328. doi: 10.1186/s12917-024-04186-6. BMC Vet Res. 2024. PMID: 39033103 Free PMC article.
-
Epidemiology, genetic diversity and evolution of pigeon circovirus.Poult Sci. 2025 Apr;104(4):104928. doi: 10.1016/j.psj.2025.104928. Epub 2025 Feb 23. Poult Sci. 2025. PMID: 40058004 Free PMC article.
-
A pan-genotypic indirect competitive ELISA for serological detection of pigeon circovirus antibodies.Front Microbiol. 2025 Jul 30;16:1612715. doi: 10.3389/fmicb.2025.1612715. eCollection 2025. Front Microbiol. 2025. PMID: 40809059 Free PMC article.
-
Rotaviruses in Pigeons With Diarrhea: Recovery of Three Complete Pigeon Rotavirus A Genomes and the First Case of Pigeon Rotavirus G in Europe.Transbound Emerg Dis. 2024 Nov 25;2024:4684235. doi: 10.1155/tbed/4684235. eCollection 2024. Transbound Emerg Dis. 2024. PMID: 40303061 Free PMC article.
References
-
- Moreira FX, et al. The use of feathers from racing pigeons for doping control purposes. J. Anal. Toxicol. 2019;43:307–315. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

