GprC of the nematode-trapping fungus Arthrobotrys flagrans activates mitochondria and reprograms fungal cells for nematode hunting
- PMID: 38877225
- PMCID: PMC11222155
- DOI: 10.1038/s41564-024-01731-9
GprC of the nematode-trapping fungus Arthrobotrys flagrans activates mitochondria and reprograms fungal cells for nematode hunting
Abstract
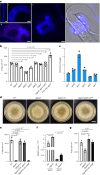
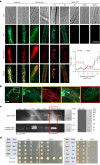
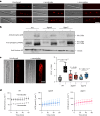
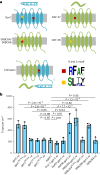
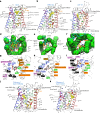
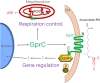
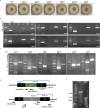
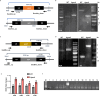
Initiation of development requires differential gene expression and metabolic adaptations. Here we show in the nematode-trapping fungus, Arthrobotrys flagrans, that both are achieved through a dual-function G-protein-coupled receptor (GPCR). A. flagrans develops adhesive traps and recognizes its prey, Caenorhabditis elegans, through nematode-specific pheromones (ascarosides). Gene-expression analyses revealed that ascarosides activate the fungal GPCR, GprC, at the plasma membrane and together with the G-protein alpha subunit GasA, reprograms the cell. However, GprC and GasA also reside in mitochondria and boost respiration. This dual localization of GprC in A. flagrans resembles the localization of the cannabinoid receptor CB1 in humans. The C. elegans ascaroside-sensing GPCR, SRBC66 and GPCRs of many fungi are also predicted for dual localization, suggesting broad evolutionary conservation. An SRBC64/66-GprC chimaeric protein was functional in A. flagrans, and C. elegans SRBC64/66 and DAF38 share ascaroside-binding sites with the fungal GprC receptor, suggesting 400-million-year convergent evolution.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
The nematode-trapping fungus Arthrobotrys oligospora detects prey pheromones via G protein-coupled receptors.Nat Microbiol. 2024 Jul;9(7):1738-1751. doi: 10.1038/s41564-024-01679-w. Epub 2024 Apr 22. Nat Microbiol. 2024. PMID: 38649409 Free PMC article.
-
The Egh16-like virulence factor TrsA of the nematode-trapping fungus Arthrobotrys flagrans facilitates intrusion into its host Caenorhabditis elegans.PLoS Pathog. 2025 Aug 25;21(8):e1013370. doi: 10.1371/journal.ppat.1013370. eCollection 2025 Aug. PLoS Pathog. 2025. PMID: 40853931 Free PMC article.
-
The cysteine-rich virulence factor NipA of Arthrobotrys flagrans interferes with cuticle integrity of Caenorhabditis elegans.Nat Commun. 2024 Jul 10;15(1):5795. doi: 10.1038/s41467-024-50096-4. Nat Commun. 2024. PMID: 38987250 Free PMC article.
-
Sensing host and environmental cues by fungal GPCRs.Curr Opin Plant Biol. 2024 Dec;82:102667. doi: 10.1016/j.pbi.2024.102667. Epub 2024 Nov 19. Curr Opin Plant Biol. 2024. PMID: 39566322 Review.
-
Ascaroside signaling in C. elegans.WormBook. 2013 Jan 18:1-22. doi: 10.1895/wormbook.1.155.1. WormBook. 2013. PMID: 23355522 Free PMC article. Review.
Cited by
-
Electron-Transferring Flavoprotein and Its Dehydrogenase Required for Fungal Pathogenicity in Arthrobotrys oligospora.Int J Mol Sci. 2024 Oct 11;25(20):10934. doi: 10.3390/ijms252010934. Int J Mol Sci. 2024. PMID: 39456717 Free PMC article.
-
AoRan1 Is Involved in Regulating Conidiation, Stress Resistance, Secondary Metabolism, and Pathogenicity in Arthrobotrys oligospora.Microorganisms. 2024 Sep 6;12(9):1853. doi: 10.3390/microorganisms12091853. Microorganisms. 2024. PMID: 39338527 Free PMC article.
-
Transcription Factors in Biocontrol Fungi.J Fungi (Basel). 2025 Mar 15;11(3):223. doi: 10.3390/jof11030223. J Fungi (Basel). 2025. PMID: 40137261 Free PMC article. Review.
-
MSB2-activated pheromone pathway regulates fungal plasma membrane integrity in response to herbicide adjuvant.Sci Adv. 2025 Feb 28;11(9):eadt8715. doi: 10.1126/sciadv.adt8715. Epub 2025 Feb 28. Sci Adv. 2025. PMID: 40020065 Free PMC article.
-
Predation by nematode-trapping fungus triggers mechanosensory-dependent quiescence in Caenorhabditis elegans.iScience. 2025 May 30;28(7):112792. doi: 10.1016/j.isci.2025.112792. eCollection 2025 Jul 18. iScience. 2025. PMID: 40585354 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

