Predicting species invasiveness with genomic data: Is genomic offset related to establishment probability?
- PMID: 38884022
- PMCID: PMC11178484
- DOI: 10.1111/eva.13709
Predicting species invasiveness with genomic data: Is genomic offset related to establishment probability?
Abstract
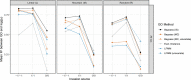
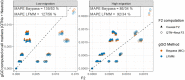
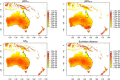
Predicting the risk of establishment and spread of populations outside their native range represents a major challenge in evolutionary biology. Various methods have recently been developed to estimate population (mal)adaptation to a new environment with genomic data via so-called Genomic Offset (GO) statistics. These approaches are particularly promising for studying invasive species but have still rarely been used in this context. Here, we evaluated the relationship between GO and the establishment probability of a population in a new environment using both in silico and empirical data. First, we designed invasion simulations to evaluate the ability to predict establishment probability of two GO computation methods (Geometric GO and Gradient Forest) under several conditions. Additionally, we aimed to evaluate the interpretability of absolute Geometric GO values, which theoretically represent the adaptive genetic distance between populations from distinct environments. Second, utilizing public empirical data from the crop pest species Bactrocera tryoni, a fruit fly native from Northern Australia, we computed GO between "source" populations and a diverse range of locations within invaded areas. This practical application of GO within the context of a biological invasion underscores its potential in providing insights and guiding recommendations for future invasion risk assessment. Overall, our results suggest that GO statistics represent good predictors of the establishment probability and may thus inform invasion risk, although the influence of several factors on prediction performance (e.g., propagule pressure or admixture) will need further investigation.
Keywords: Bactrocera tryoni; GEA; biological invasions; genomic offset; local adaptation.
© 2024 The Author(s). Evolutionary Applications published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors have no conflict of interest to disclose.
Figures


Similar articles
-
Genomic signals of local adaptation across climatically heterogenous habitats in an invasive tropical fruit fly (Bactrocera tryoni).Heredity (Edinb). 2024 Jan;132(1):18-29. doi: 10.1038/s41437-023-00657-y. Epub 2023 Oct 30. Heredity (Edinb). 2024. PMID: 37903919 Free PMC article.
-
Parasitoids of Queensland Fruit Fly Bactrocera tryoni in Australia and Prospects for Improved Biological Control.Insects. 2012 Oct 22;3(4):1056-83. doi: 10.3390/insects3041056. Insects. 2012. PMID: 26466726 Free PMC article. Review.
-
Incorporating adaptive genomic variation into predictive models for invasion risk assessment.Environ Sci Ecotechnol. 2023 Jul 11;18:100299. doi: 10.1016/j.ese.2023.100299. eCollection 2024 Mar. Environ Sci Ecotechnol. 2023. PMID: 37701243 Free PMC article.
-
Multiple introductions, admixture and bridgehead invasion characterize the introduction history of Ambrosia artemisiifolia in Europe and Australia.Mol Ecol. 2017 Oct;26(20):5421-5434. doi: 10.1111/mec.14293. Epub 2017 Sep 11. Mol Ecol. 2017. PMID: 28802079
-
The evolutionary dynamics of biological invasions: A multi-approach perspective.Evol Appl. 2021 Mar 30;14(6):1463-1484. doi: 10.1111/eva.13215. eCollection 2021 Jun. Evol Appl. 2021. PMID: 34178098 Free PMC article. Review.
Cited by
-
Genomic Reconstruction of the Successful Establishment of a Feralized Bovine Population on the Subantarctic Island of Amsterdam.Mol Biol Evol. 2024 Jul 3;41(7):msae121. doi: 10.1093/molbev/msae121. Mol Biol Evol. 2024. PMID: 38889245 Free PMC article.
References
-
- Aravind, N. , Shaanker, M. , Bhat, P. , Charles, B. , Uma Shaanker, R. , Shah, M. , & Ravikanth, G. (2022). Niche shift in invasive species: Is it a case of “home away from home” or finding a “new home”? Biodiversity and Conservation, 31, 1–14.
-
- Archambeau, J. (2022). Understanding the origin and predicting adaptive genetic variation at large scale in the genomic era: A case study in maritime pine . These de doctorat, Bordeaux.
-
- Barrett, R. D. H. , & Schluter, D. (2008). Adaptation from standing genetic variation. Trends in Ecology & Evolution, 23(1), 38–44. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

